+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4007 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of iFuse PFV glycoprotein by cryo-electron tomography | |||||||||
 Map data Map data | PFV iFuse Glycoprotein | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Human spumaretrovirus Human spumaretrovirus | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 28.0 Å | |||||||||
 Authors Authors | Effantin G | |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2016 Journal: PLoS Pathog / Year: 2016Title: Cryo-electron Microscopy Structure of the Native Prototype Foamy Virus Glycoprotein and Virus Architecture. Authors: Grégory Effantin / Leandro F Estrozi / Nick Aschman / Patricia Renesto / Nicole Stanke / Dirk Lindemann / Guy Schoehn / Winfried Weissenhorn /   Abstract: Foamy viruses (FV) belong to the genus Spumavirus, which forms a distinct lineage in the Retroviridae family. Although the infection in natural hosts and zoonotic transmission to humans is ...Foamy viruses (FV) belong to the genus Spumavirus, which forms a distinct lineage in the Retroviridae family. Although the infection in natural hosts and zoonotic transmission to humans is asymptomatic, FVs can replicate well in human cells making it an attractive gene therapy vector candidate. Here we present cryo-electron microscopy and (cryo-)electron tomography ultrastructural data on purified prototype FV (PFV) and PFV infected cells. Mature PFV particles have a distinct morphology with a capsid of constant dimension as well as a less ordered shell of density between the capsid and the membrane likely formed by the Gag N-terminal domain and the cytoplasmic part of the Env leader peptide gp18LP. The viral membrane contains trimeric Env glycoproteins partly arranged in interlocked hexagonal assemblies. In situ 3D reconstruction by subtomogram averaging of wild type Env and of a Env gp48TM- gp80SU cleavage site mutant showed a similar spike architecture as well as stabilization of the hexagonal lattice by clear connections between lower densities of neighboring trimers. Cryo-EM was employed to obtain a 9 Å resolution map of the glycoprotein in its pre-fusion state, which revealed extensive trimer interactions by the receptor binding subunit gp80SU at the top of the spike and three central helices derived from the fusion protein subunit gp48TM. The lower part of Env, presumably composed of interlaced parts of gp48TM, gp80SU and gp18LP anchors the spike at the membrane. We propose that the gp48TM density continues into three central transmembrane helices, which interact with three outer transmembrane helices derived from gp18LP. Our ultrastructural data and 9 Å resolution glycoprotein structure provide important new insights into the molecular architecture of PFV and its distinct evolutionary relationship with other members of the Retroviridae. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4007.map.gz emd_4007.map.gz | 1.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4007-v30.xml emd-4007-v30.xml emd-4007.xml emd-4007.xml | 8.4 KB 8.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4007.png emd_4007.png | 96.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4007 http://ftp.pdbj.org/pub/emdb/structures/EMD-4007 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4007 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4007 | HTTPS FTP |
-Validation report
| Summary document |  emd_4007_validation.pdf.gz emd_4007_validation.pdf.gz | 77.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_4007_full_validation.pdf.gz emd_4007_full_validation.pdf.gz | 76.6 KB | Display | |
| Data in XML |  emd_4007_validation.xml.gz emd_4007_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4007 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4007 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4007 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4007 | HTTPS FTP |
-Related structure data
| Related structure data | 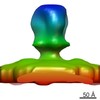 4006C  4008C  4010C  4011C  4012C  4013C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4007.map.gz / Format: CCP4 / Size: 1.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4007.map.gz / Format: CCP4 / Size: 1.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PFV iFuse Glycoprotein | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human spumaretrovirus
| Entire | Name:  Human spumaretrovirus Human spumaretrovirus |
|---|---|
| Components |
|
-Supramolecule #1: Human spumaretrovirus
| Supramolecule | Name: Human spumaretrovirus / type: virus / ID: 1 / Parent: 0 / Details: Mutant in the Env glycoprotein / NCBI-ID: 11963 / Sci species name: Human spumaretrovirus / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host system | Organism:  Homo sapiens (human) / Recombinant plasmid: pcoPG4, pcoPE32, pcoPP Homo sapiens (human) / Recombinant plasmid: pcoPG4, pcoPE32, pcoPP |
| Molecular weight | Theoretical: 330 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
| Details | The iFuse mutant is a variant of Env where the furine cleavage site between the SUrface (gp80 SU ) and TransMembrane (gp48 TM ) domains of Env has been mutated. This results in a partially processed glycoprotein as the cleavage between the LP and SU domains is preserved. Particles are released at nearly wild type level from cells but are non-infectious |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: FEI EAGLE (4k x 4k) / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 28.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: PEET / Number subtomograms used: 3000 |
|---|---|
| Extraction | Number tomograms: 7 / Number images used: 3930 |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller








