[English] 日本語
 Yorodumi
Yorodumi- EMDB-2394: Architecture of an RNA polymerase II transcription pre-initiation... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2394 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Architecture of an RNA polymerase II transcription pre-initiation complex | |||||||||
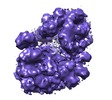
 Map data Map data | Reconstruction of RNA polymerase II transcription pre-initiation complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA Polymerase II / Transcription / Transcription Initiation Factors / Yeast / TFIIA / TFIIB / TBP / TFIIE / TFIIF / TFIIH / TFIIS | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 16.0 Å | |||||||||
 Authors Authors | Murakami K / Elmlund H / Kalisman N / Bushnell DA / Adams CM / Azubel M / Elmlund D / Levi-Kalisman Y / Liu X / Levitt M / Kornberg RD | |||||||||
 Citation Citation |  Journal: Science / Year: 2013 Journal: Science / Year: 2013Title: Architecture of an RNA polymerase II transcription pre-initiation complex. Authors: Kenji Murakami / Hans Elmlund / Nir Kalisman / David A Bushnell / Christopher M Adams / Maia Azubel / Dominika Elmlund / Yael Levi-Kalisman / Xin Liu / Brian J Gibbons / Michael Levitt / Roger D Kornberg /  Abstract: The protein density and arrangement of subunits of a complete, 32-protein, RNA polymerase II (pol II) transcription pre-initiation complex (PIC) were determined by means of cryogenic electron ...The protein density and arrangement of subunits of a complete, 32-protein, RNA polymerase II (pol II) transcription pre-initiation complex (PIC) were determined by means of cryogenic electron microscopy and a combination of chemical cross-linking and mass spectrometry. The PIC showed a marked division in two parts, one containing all the general transcription factors (GTFs) and the other pol II. Promoter DNA was associated only with the GTFs, suspended above the pol II cleft and not in contact with pol II. This structural principle of the PIC underlies its conversion to a transcriptionally active state; the PIC is poised for the formation of a transcription bubble and descent of the DNA into the pol II cleft. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2394.map.gz emd_2394.map.gz | 28.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2394-v30.xml emd-2394-v30.xml emd-2394.xml emd-2394.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2394.png EMD-2394.png | 172.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2394 http://ftp.pdbj.org/pub/emdb/structures/EMD-2394 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2394 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2394 | HTTPS FTP |
-Validation report
| Summary document |  emd_2394_validation.pdf.gz emd_2394_validation.pdf.gz | 202.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2394_full_validation.pdf.gz emd_2394_full_validation.pdf.gz | 201.4 KB | Display | |
| Data in XML |  emd_2394_validation.xml.gz emd_2394_validation.xml.gz | 6.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2394 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2394 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2394 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2394 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2394.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2394.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of RNA polymerase II transcription pre-initiation complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.85 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : RNA polymerase II transcription pre-initiation complex
| Entire | Name: RNA polymerase II transcription pre-initiation complex |
|---|---|
| Components |
|
-Supramolecule #1000: RNA polymerase II transcription pre-initiation complex
| Supramolecule | Name: RNA polymerase II transcription pre-initiation complex type: sample / ID: 1000 / Number unique components: 32 |
|---|---|
| Molecular weight | Experimental: 1.5 MDa / Theoretical: 1.5 MDa |
-Macromolecule #1: RNA polymerase II transcription pre-initiation complex
| Macromolecule | Name: RNA polymerase II transcription pre-initiation complex type: protein_or_peptide / ID: 1 / Name.synonym: PIC / Details: including HIS4(-81/+1) template DNA / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 1.5 MDa / Theoretical: 1.5 MDa |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.6 Details: 20mM HEPES, 5mM DTT, 2mM magnesium acetate, 30 mM potassium acetate |
| Staining | Type: NEGATIVE Details: 20 ul of fixed sample was dialyzed for 2 hours to remove glycerol. 3 ul were transferred to a grid and flash frozen in liquid ethane with a Vitrobot. |
| Grid | Details: Quantifoil was glow-discharged using a Harrick Plasma Cleaner |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 120 K / Instrument: FEI VITROBOT MARK III / Method: blot time 3 sec, waiting 10sec, force 9 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Min: 80 K / Max: 105 K / Average: 100 K |
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 81,000 times magnification |
| Date | Jun 25, 2010 |
| Image recording | Category: CCD / Film or detector model: GENERIC GATAN (4k x 4k) / Average electron dose: 15 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 81000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.26 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | PIC particles were windowed semi-automatically using the EMAN program Boxer |
|---|---|
| CTF correction | Details: frame |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 16.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SIMPLE / Number images used: 433916 |
 Movie
Movie Controller
Controller