+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1792 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
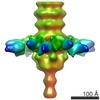
| Title | EM map of TP901-1 BppU-BppL complex | |||||||||
 Map data Map data | EM map of TP9101-1 BppU-BppL complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Bacteriophage / TP9101-1 / BppU / BppL / ORF 48 / ORF 49 Bacteriophage / TP9101-1 / BppU / BppL / ORF 48 / ORF 49 | |||||||||
| Biological species |  Lactococcus phage TP901-1 (virus) Lactococcus phage TP901-1 (virus) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  negative staining / Resolution: 24.8 Å negative staining / Resolution: 24.8 Å | |||||||||
 Authors Authors | Bron P | |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2010 Journal: J Biol Chem / Year: 2010Title: Structure and molecular assignment of lactococcal phage TP901-1 baseplate. Authors: Cecilia Bebeacua / Patrick Bron / Livia Lai / Christina Skovgaard Vegge / Lone Brøndsted / Silvia Spinelli / Valérie Campanacci / David Veesler / Marin van Heel / Christian Cambillau /  Abstract: P335 lactococcal phages infect the gram(+) bacterium Lactococcus lactis using a large multiprotein complex located at the distal part of the tail and termed baseplate (BP). The BP harbors the ...P335 lactococcal phages infect the gram(+) bacterium Lactococcus lactis using a large multiprotein complex located at the distal part of the tail and termed baseplate (BP). The BP harbors the receptor-binding proteins (RBPs), which allow the specific recognition of saccharidic receptors localized on the host cell surface. We report here the electron microscopic structure of the phage TP901-1 wild-type BP as well as those of two mutants bppL (-) and bppU(-), lacking BppL (the RBPs) or both peripheral BP components (BppL and BppU), respectively. We also achieved an electron microscopic reconstruction of a partial BP complex, formed by BppU and BppL. This complex exhibits a tripod shape and is composed of nine BppLs and three BppUs. These structures, combined with light-scattering measurements, led us to propose that the TP901-1 BP harbors six tripods at its periphery, located around the central tube formed by ORF46 (Dit) hexamers, at its proximal end, and a ORF47 (Tal) trimer at its distal extremity. A total of 54 BppLs (18 RBPs) are thus available to mediate host anchoring with a large apparent avidity. TP901-1 BP exhibits an infection-ready conformation and differs strikingly from the lactococcal phage p2 BP, bearing only 6 RBPs, and which needs a conformational change to reach its activated state. The comparison of several Siphoviridae structures uncovers a close organization of their central BP core whereas striking differences occur at the periphery, leading to diverse mechanisms of host recognition. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1792.map.gz emd_1792.map.gz | 2.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1792-v30.xml emd-1792-v30.xml emd-1792.xml emd-1792.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-1792.png EMD-1792.png | 46 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1792 http://ftp.pdbj.org/pub/emdb/structures/EMD-1792 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1792 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1792 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1792.map.gz / Format: CCP4 / Size: 2.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1792.map.gz / Format: CCP4 / Size: 2.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM map of TP9101-1 BppU-BppL complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : BppU/BppL complex from Lactococcal Phage TP901-1
| Entire | Name: BppU/BppL complex from Lactococcal Phage TP901-1 |
|---|---|
| Components |
|
-Supramolecule #1000: BppU/BppL complex from Lactococcal Phage TP901-1
| Supramolecule | Name: BppU/BppL complex from Lactococcal Phage TP901-1 / type: sample / ID: 1000 Oligomeric state: one trimer of BppU and three trimers of BppL Number unique components: 2 |
|---|
-Macromolecule #1: BppU
| Macromolecule | Name: BppU / type: protein_or_peptide / ID: 1 / Name.synonym: ORF48 / Number of copies: 1 / Oligomeric state: Monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Lactococcus phage TP901-1 (virus) Lactococcus phage TP901-1 (virus) |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
-Macromolecule #2: BppL
| Macromolecule | Name: BppL / type: protein_or_peptide / ID: 2 / Name.synonym: ORF49 / Number of copies: 3 / Oligomeric state: Trimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Lactococcus phage TP901-1 (virus) Lactococcus phage TP901-1 (virus) |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
-Experimental details
-Structure determination
| Method |  negative staining negative staining |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.05 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 10 mM Hepes, 150 mM NaCl, 1 mM PMSF |
| Staining | Type: NEGATIVE Details: Grids with adsorbed protein floated on 1% uranyl acetate for 1 min. |
| Grid | Details: 400 mesh cupper griid |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2200FS |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 50000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.0 mm / Nominal defocus max: 1.0 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 50000 Bright-field microscopy / Cs: 2.0 mm / Nominal defocus max: 1.0 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 50000 |
| Specialist optics | Energy filter - Name: Omega filter / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: JEOL |
| Alignment procedure | Legacy - Astigmatism: Astigmatism was corrected at 150,000 times magnification |
| Details | Low-dose imaging |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 10 µm / Number real images: 8 / Average electron dose: 18 e/Å2 / Bits/pixel: 8 |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic ) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 24.8 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: IMAGIC V / Number images used: 8314 ) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 24.8 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: IMAGIC V / Number images used: 8314 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Details | The trimers were separately fitted by manual docking and refined using the docking program integrated into Chimera |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller