+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1785 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The Structure of a COPII Tubule | |||||||||
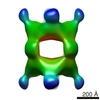
 Map data Map data | Isosurface contour map of Subvolume averaged Cuboctahedron cage. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Secretory pathway / Cryo-electron microscopy / Cryo-electron tomography / COPII / cargo / Subvolume averaging | |||||||||
| Function / homology | intracellular protein transport / WD40 repeat Function and homology information Function and homology information | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 85.0 Å | |||||||||
 Authors Authors | ODonnell J / Maddox K / Stagg S | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2011 Journal: J Struct Biol / Year: 2011Title: The structure of a COPII tubule. Authors: Jason O'Donnell / Kerry Maddox / Scott Stagg /  Abstract: Nearly a third of all eukaryotic proteins are transported from the ER to the Golgi apparatus through the secretory pathway using COPII coated vesicles. Evidence suggests that this transport occurs ...Nearly a third of all eukaryotic proteins are transported from the ER to the Golgi apparatus through the secretory pathway using COPII coated vesicles. Evidence suggests that this transport occurs via 500-900 Å vesicles that bud from the ER membrane. It has been shown that procollagen molecules utilize the COPII proteins for transport, but it is unclear how the COPII coat can accommodate these ∼3000 Å long molecules. We now present a cryogenic electron tomographic reconstruction of a Sec13/31 tubule that is approximately 3300 Å long containing a hollow cylindrical interior that is 300 Å in diameter, dimensions that are consistent with those that are required to encapsulate a procollagen molecule wrapped in a membrane and accessory COPII components. This structure suggests a novel mechanism that the COPII coat may employ to transport elongated cargo. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1785.map.gz emd_1785.map.gz | 2.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1785-v30.xml emd-1785-v30.xml emd-1785.xml emd-1785.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| Images |  1785.jpg 1785.jpg | 50.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1785 http://ftp.pdbj.org/pub/emdb/structures/EMD-1785 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1785 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1785 | HTTPS FTP |
-Validation report
| Summary document |  emd_1785_validation.pdf.gz emd_1785_validation.pdf.gz | 193.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1785_full_validation.pdf.gz emd_1785_full_validation.pdf.gz | 192.8 KB | Display | |
| Data in XML |  emd_1785_validation.xml.gz emd_1785_validation.xml.gz | 6.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1785 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1785 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1785 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1785 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1785.map.gz / Format: CCP4 / Size: 26.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1785.map.gz / Format: CCP4 / Size: 26.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Isosurface contour map of Subvolume averaged Cuboctahedron cage. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 9.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Oligomeric assembly of Sec13-31 proteins
| Entire | Name: Oligomeric assembly of Sec13-31 proteins |
|---|---|
| Components |
|
-Supramolecule #1000: Oligomeric assembly of Sec13-31 proteins
| Supramolecule | Name: Oligomeric assembly of Sec13-31 proteins / type: sample / ID: 1000 / Oligomeric state: 24-mer made by Interlocked Cuboctahedrons / Number unique components: 2 |
|---|---|
| Molecular weight | Theoretical: 8.14 MDa |
-Macromolecule #1: SEC13R
| Macromolecule | Name: SEC13R / type: protein_or_peptide / ID: 1 / Name.synonym: Sec13 / Number of copies: 12 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Cell: Sf9 Homo sapiens (human) / synonym: Human / Cell: Sf9 |
| Recombinant expression | Organism:  |
| Sequence | GO: intracellular protein transport / InterPro: WD40 repeat |
-Macromolecule #2: SEC31L1
| Macromolecule | Name: SEC31L1 / type: protein_or_peptide / ID: 2 / Name.synonym: Sec31 / Number of copies: 12 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Cell: Sf9 Homo sapiens (human) / synonym: Human / Cell: Sf9 |
| Recombinant expression | Organism:  |
| Sequence | GO: intracellular protein transport / InterPro: WD40 repeat |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 20 mM Tris-Cl, pH 7.5, 700 mM KOAc, 1mM MgOAc, 10mM DTT |
|---|---|
| Grid | Details: Quantifoil 2/2, 400 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 93 K / Instrument: OTHER Details: Vitrification instrument: FEI Vitrobot. Grid plasma cleaned for 30s with Gatan Solarus 950 plasma cleaner using a ratio of 75-25 of Argon-Oxygen. Method: Temperature of chamber was 4 degrees C. 0 seconds drain time. Single blot. 0 mm offset. 3 ul sample applied to grid. Blot for 2.5 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM300FEG/T |
|---|---|
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F415 (4k x 4k) / Digitization - Sampling interval: 15 µm / Number real images: 64 |
| Electron beam | Acceleration voltage: 300 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 24000 |
| Sample stage | Specimen holder: Side entry liquid nitrogen-cooled cryo specimen holder. Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Max angle: 64 ° |
- Image processing
Image processing
| Details | The Subvolume motif was selected using the program PROTOMO. Average number of tilts used in the 3D reconstructions: 250. Average tomographic tilt angle increment: 2. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 85.0 Å / Resolution method: OTHER / Software - Name: PROTOMO |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera