+ Open data
Open data
- Basic information
Basic information
| Entry | 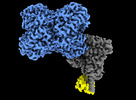 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
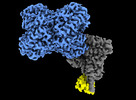
| Title | Structure of Trypanosoma docking complex | |||||||||
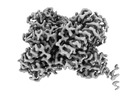
 Map data Map data | This is main composite map, generated from two component maps. Component 1 - MDH_flipped.mrc and Component 2 - PEX_fliiped.mrc, which are provided as additional maps. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Peroxisomal transport / Import / PEX /  TRANSPORT PROTEIN / TRANSPORT PROTEIN /  PROTEIN TRANSPORT PROTEIN TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein import into peroxisome matrix, docking /  malate dehydrogenase / L-malate dehydrogenase activity / peroxisomal membrane / carboxylic acid metabolic process / malate dehydrogenase / L-malate dehydrogenase activity / peroxisomal membrane / carboxylic acid metabolic process /  tricarboxylic acid cycle / tricarboxylic acid cycle /  peroxisome peroxisomeSimilarity search - Function | |||||||||
| Biological species |   Trypanosoma cruzi (eukaryote) / Trypanosoma cruzi (eukaryote) /   Trypanosoma cruzi strain CL Brener (eukaryote) / Trypanosoma cruzi strain CL Brener (eukaryote) /  | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.5 Å cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Sonani RR / Artur B / Jemiola-Rzeminska M / Lipinski O / Patel SN / Sood T / Dubin G | |||||||||
| Funding support |  Poland, 2 items Poland, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of Trypanosoma docking complex Authors: Sonani RR / Dubin G | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40056.map.gz emd_40056.map.gz | 3.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40056-v30.xml emd-40056-v30.xml emd-40056.xml emd-40056.xml | 23 KB 23 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40056.png emd_40056.png | 85.7 KB | ||
| Filedesc metadata |  emd-40056.cif.gz emd-40056.cif.gz | 6.5 KB | ||
| Others |  emd_40056_additional_1.map.gz emd_40056_additional_1.map.gz emd_40056_additional_2.map.gz emd_40056_additional_2.map.gz emd_40056_additional_3.map.gz emd_40056_additional_3.map.gz emd_40056_half_map_1.map.gz emd_40056_half_map_1.map.gz emd_40056_half_map_2.map.gz emd_40056_half_map_2.map.gz | 14.5 MB 110.4 MB 200.6 MB 200.7 MB 200.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40056 http://ftp.pdbj.org/pub/emdb/structures/EMD-40056 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40056 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40056 | HTTPS FTP |
-Related structure data
| Related structure data |  8gi0MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40056.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40056.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is main composite map, generated from two component maps. Component 1 - MDH_flipped.mrc and Component 2 - PEX_fliiped.mrc, which are provided as additional maps. | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Component 1 - MDH of coposite map. Separately...
| File | emd_40056_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Component 1 - MDH of coposite map. Separately depositied to EMBD, EMD-40053 | ||||||||||||
| Projections & Slices |
| ||||||||||||
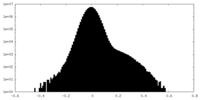
| Density Histograms |
-Additional map: #1
| File | emd_40056_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
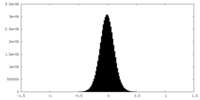
| Density Histograms |
-Additional map: #2
| File | emd_40056_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
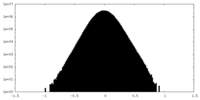
| Density Histograms |
-Half map: #1
| File | emd_40056_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
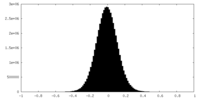
| Density Histograms |
-Half map: #2
| File | emd_40056_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Docking complex (MDH)4-(Pex5)1-(Pex14)1 of Trypanosoma Pex import
| Entire | Name: Docking complex (MDH)4-(Pex5)1-(Pex14)1 of Trypanosoma Pex import |
|---|---|
| Components |
|
-Supramolecule #1: Docking complex (MDH)4-(Pex5)1-(Pex14)1 of Trypanosoma Pex import
| Supramolecule | Name: Docking complex (MDH)4-(Pex5)1-(Pex14)1 of Trypanosoma Pex import type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Trypanosoma cruzi (eukaryote) Trypanosoma cruzi (eukaryote) |
-Macromolecule #1: Peroxisomal membrane protein PEX14
| Macromolecule | Name: Peroxisomal membrane protein PEX14 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Trypanosoma cruzi (eukaryote) Trypanosoma cruzi (eukaryote) |
| Molecular weight | Theoretical: 40.507551 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: MNTATTVGVT GDGDARQRNS PEESEKRKRV SSAVQFLHDS RVKITPAANK IQFLKSKGLT TEEVCEAFEK AGQTIPLDEI KKIMNKPSF GQLGSGVVNN NLAPGHVGSD ATYVLRQHPF MPHAGPLYTQ QPPPFPQTPQ GAKETDWRDV IIGVGAALIS G FAAFKAFQ ...String: MNTATTVGVT GDGDARQRNS PEESEKRKRV SSAVQFLHDS RVKITPAANK IQFLKSKGLT TEEVCEAFEK AGQTIPLDEI KKIMNKPSF GQLGSGVVNN NLAPGHVGSD ATYVLRQHPF MPHAGPLYTQ QPPPFPQTPQ GAKETDWRDV IIGVGAALIS G FAAFKAFQ IYSPYEIRRK DERTKKGSRM ESQRRGKRHE GSVSPDRGAE RPVTSLHTMA PPMPATPTPL PTATAANAAE ST EREIQKL QSELKETQEA LEKEKRSKAE ISVSLGKLRG QINALSRTND NLESRIKLLQ EEVDKANSEA NKRKELDASF TGA ADGVPK SQSEDAIQPN GPPLLTPKIL ADTVGCNSAA LNELPVLTAV EFVPEE UniProtKB: Peroxisomal membrane protein PEX14 |
-Macromolecule #2: malate dehydrogenase
| Macromolecule | Name: malate dehydrogenase / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Trypanosoma cruzi strain CL Brener (eukaryote) Trypanosoma cruzi strain CL Brener (eukaryote) |
| Molecular weight | Theoretical: 34.106832 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: MVNVAVIGAA GGIGQSLSLL LLRELPFGST LSLYDVVGAP GVAADLSHID RAGITVKHAA GKLPPVPRDP ALTELAEGVD VFVIVAGVP RKPGMTRDDL FNVNAGIVMD LVLTCASVSP NACFCIVTNP VNSTTPIAAQ TLRKIGVYNK NKLLGVSLLD G LRATRFIN ...String: MVNVAVIGAA GGIGQSLSLL LLRELPFGST LSLYDVVGAP GVAADLSHID RAGITVKHAA GKLPPVPRDP ALTELAEGVD VFVIVAGVP RKPGMTRDDL FNVNAGIVMD LVLTCASVSP NACFCIVTNP VNSTTPIAAQ TLRKIGVYNK NKLLGVSLLD G LRATRFIN NARHPLVVPY VPVVGGHSDV TIVPLYSQIP GPLPDESTLK EIRKRVQVAG TEVVKAKAGR GSATLSMAEA GA RFTMHVV KALMGLDTPM VYAYVDTDGE HECPFLAMPV VLGKNGIERR LPIGPITTVE KEMLEEAVGV VKKNIAKGET FAR SKL UniProtKB:  malate dehydrogenase malate dehydrogenase |
-Macromolecule #3: Peroxisome targeting signal 1 receptor
| Macromolecule | Name: Peroxisome targeting signal 1 receptor / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 74.268422 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: MDCSTGAAIG QQFAKDAFHM HGGVGVGPTG NSEHDVLMNE MMMVQTPTGP AGEWTHQFAA YQGQQQQQQQ QHPQELAMRH QQNDAFMLR QQQEMEEAFC TFCTTHPHSH AHSHQPQGLV GPAMMGPQIM PPMMFGPGTG GFMMGAPPMM PYASMKFAGD A AMAAANNT ...String: MDCSTGAAIG QQFAKDAFHM HGGVGVGPTG NSEHDVLMNE MMMVQTPTGP AGEWTHQFAA YQGQQQQQQQ QHPQELAMRH QQNDAFMLR QQQEMEEAFC TFCTTHPHSH AHSHQPQGLV GPAMMGPQIM PPMMFGPGTG GFMMGAPPMM PYASMKFAGD A AMAAANNT NMTQGATATS TTSVQQELQQ QSSDNGWVEK LRDAEWAQDY SDAQVFTLEG QSEQTMEEHA KNSEFYQFMD KI RSKELLI DEETGQLVQG PGPDPDAPED AEYLKEWAAA EGLNMPPGFF EHMMQRPQGN NEQAEGRLFD GSNDALMDDG ALD NAADVE EWVREYAEAQ EQLQRVQNET NYPFEPNNPY MYHDKPMEEG IAMLQLANMA EAALAFEAVC QKEPENVEAW RRLG TTQAE NEKDCLAIIA LNHARMLDPK DIAVHAALAV SHTNEHNVGA ALQSLRSWLL SQPQYEHLGL VDLREVAADE GLDEV PEEN YFFAAPSEYR DCCTLLYAAV EMNPNDPQLH ASLGVLHNLS HRFDEAAKNF RRAVELRPDD AHMWNKLGAT LANGNR PQE ALEAYNRALD INPGYVRVMY NMAVSYSNMA QYPLAAKHIT RAIALQAGGT NPQGEGSRIA TRGLWDLLRM TLNLMDR SD LVEASWQQDL TPFLREFGLE EMAV UniProtKB: Peroxisome targeting signal 1 receptor |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.9 µm Bright-field microscopy / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.9 µm |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 42.25 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 482240 |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X