[English] 日本語
 Yorodumi
Yorodumi- EMDB-38261: Cryo-EM structure of Glutamate dehydrogenase from Thermococcus pr... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Glutamate dehydrogenase from Thermococcus profundus incorporating NADPH and AKG in the steady stage of reaction | |||||||||||||||||||||||||||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||||||||||||||||||||
 Keywords Keywords |  Complex / Complex /  Coenzyme / Coenzyme /  NADPH / NADPH /  2-oxoglutarate / 2-oxoglutarate /  OXIDOREDUCTASE OXIDOREDUCTASE | |||||||||||||||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationglutamate dehydrogenase [NAD(P)+] /  glutamate dehydrogenase (NADP+) activity / glutamate dehydrogenase (NAD+) activity / amino acid metabolic process glutamate dehydrogenase (NADP+) activity / glutamate dehydrogenase (NAD+) activity / amino acid metabolic processSimilarity search - Function | |||||||||||||||||||||||||||||||||||||||||||||
| Biological species |    Thermococcus profundus (archaea) Thermococcus profundus (archaea) | |||||||||||||||||||||||||||||||||||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.83 Å cryo EM / Resolution: 2.83 Å | |||||||||||||||||||||||||||||||||||||||||||||
 Authors Authors | Wakabayashi T / Oide M / Nakasako M | |||||||||||||||||||||||||||||||||||||||||||||
| Funding support |  Japan, 14 items Japan, 14 items
| |||||||||||||||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: CryoEM-sampling of metastable conformations appearing in cofactor-ligand association and catalysis of glutamate dehydrogenase Authors: Wakabayashi T / Oide M / Nakasako M | |||||||||||||||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38261.map.gz emd_38261.map.gz | 26.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38261-v30.xml emd-38261-v30.xml emd-38261.xml emd-38261.xml | 20.9 KB 20.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38261_fsc.xml emd_38261_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_38261.png emd_38261.png | 151 KB | ||
| Masks |  emd_38261_msk_1.map emd_38261_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-38261.cif.gz emd-38261.cif.gz | 6.5 KB | ||
| Others |  emd_38261_half_map_1.map.gz emd_38261_half_map_1.map.gz emd_38261_half_map_2.map.gz emd_38261_half_map_2.map.gz | 60 MB 60 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38261 http://ftp.pdbj.org/pub/emdb/structures/EMD-38261 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38261 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38261 | HTTPS FTP |
-Related structure data
| Related structure data |  8xd0MC  8xcoC  8xcpC  8xcqC  8xcrC  8xcsC  8xctC  8xcuC  8xcvC  8xcwC  8xcxC  8xcyC  8xczC  8xd1C  8xd2C  8xd3C  8xd4C  8xd5C  8xd6C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_38261.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38261.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.752 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_38261_msk_1.map emd_38261_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
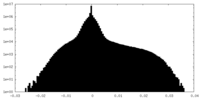
| Density Histograms |
-Half map: #2
| File | emd_38261_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_38261_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Hexamer of glutamate dehydrogenase in the presence of NADP and gl...
| Entire | Name: Hexamer of glutamate dehydrogenase in the presence of NADP and glutamate |
|---|---|
| Components |
|
-Supramolecule #1: Hexamer of glutamate dehydrogenase in the presence of NADP and gl...
| Supramolecule | Name: Hexamer of glutamate dehydrogenase in the presence of NADP and glutamate type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:    Thermococcus profundus (archaea) Thermococcus profundus (archaea) |
| Molecular weight | Theoretical: 280 KDa |
-Macromolecule #1: Glutamate dehydrogenase
| Macromolecule | Name: Glutamate dehydrogenase / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: glutamate dehydrogenase [NAD(P)+] |
|---|---|
| Source (natural) | Organism:    Thermococcus profundus (archaea) Thermococcus profundus (archaea) |
| Molecular weight | Theoretical: 46.758477 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MVEIDPFEMA VKQLERAAQY MDISEEALEW LKKPMRIVEV SVPIEMDDGS VKVFTGFRVQ HNWARGPTKG GIRWHPAETL STVKALATW MTWKVAVVDL PYGGGKGGII VNPKELSERE QERLARAYIR AVYDVIGPWT DIPAPDVYTN PKIMGWMMDE Y ETIMRRKG ...String: MVEIDPFEMA VKQLERAAQY MDISEEALEW LKKPMRIVEV SVPIEMDDGS VKVFTGFRVQ HNWARGPTKG GIRWHPAETL STVKALATW MTWKVAVVDL PYGGGKGGII VNPKELSERE QERLARAYIR AVYDVIGPWT DIPAPDVYTN PKIMGWMMDE Y ETIMRRKG PAFGVITGKP LSIGGSLGRG TATAQGAIFT IREAAKALGI DLKGKKIAVQ GYGNAGYYTA KLAKEQLGMT VV AVSDSRG GIYNPDGLDP DEVLKWKREH GSVKDFPGAT NITNEELLEL EVDVLAPAAI EEVITEKNAD NIKAKIVAEV ANG PVTPEA DDILREKGIL QIPDFLCNAG GVTVSYFEWV QNINGYYWTE EEVREKLDKK MTKAFWEVYN THKDKNIHMR DAAY VVAVS RVYQAMKDRG WVKK UniProtKB:  Glutamate dehydrogenase Glutamate dehydrogenase |
-Macromolecule #2: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
| Macromolecule | Name: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE type: ligand / ID: 2 / Number of copies: 1 / Formula: NDP |
|---|---|
| Molecular weight | Theoretical: 745.421 Da |
| Chemical component information |  ChemComp-NDP: |
-Macromolecule #3: 2-OXOGLUTARIC ACID
| Macromolecule | Name: 2-OXOGLUTARIC ACID / type: ligand / ID: 3 / Number of copies: 1 / Formula: AKG |
|---|---|
| Molecular weight | Theoretical: 146.098 Da |
| Chemical component information |  ChemComp-AKG: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 14 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.8 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV Details: The sample solution kept at room temperature was flash-frozen 1-h after mixing GDH, NADP and glutamate solutions.. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 47.0 µm / Nominal defocus min: 0.4 µm Bright-field microscopy / Nominal defocus max: 47.0 µm / Nominal defocus min: 0.4 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Details | Grid information as following: Company/model: Quantifoil Cu 1.2/1.3 Material:Cu Grid mesh: 200 |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 7075 / Average electron dose: 1.0 e/Å2 |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8xd0: |
 Movie
Movie Controller
Controller



















 Z
Z Y
Y X
X