[English] 日本語
 Yorodumi
Yorodumi- EMDB-3320: CryoEM structure of the ATP-treated CMG replicative helicase (com... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3320 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of the ATP-treated CMG replicative helicase (compact state) | |||||||||
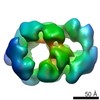
 Map data Map data | CMG treated with ATP in a tight Mcm5-2 AAA+ configuration | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cdc45 / GINS / MCM / CMG / helicase / DNA replication | |||||||||
| Function / homology |  Function and homology information Function and homology informationUnwinding of DNA / Switching of origins to a post-replicative state / Assembly of the pre-replicative complex / DNA endoreduplication / Activation of ATR in response to replication stress / Activation of the pre-replicative complex / eggshell chorion gene amplification / Orc1 removal from chromatin / DNA amplification / GINS complex ...Unwinding of DNA / Switching of origins to a post-replicative state / Assembly of the pre-replicative complex / DNA endoreduplication / Activation of ATR in response to replication stress / Activation of the pre-replicative complex / eggshell chorion gene amplification / Orc1 removal from chromatin / DNA amplification / GINS complex / DNA strand elongation involved in mitotic DNA replication / mitotic DNA replication preinitiation complex assembly / resolution of meiotic recombination intermediates / premeiotic DNA replication / mitotic DNA replication / CMG complex / MCM complex / DNA replication preinitiation complex / double-strand break repair via break-induced replication / mitotic DNA replication initiation / chromosome condensation / DNA duplex unwinding / DNA strand elongation involved in DNA replication / DNA unwinding involved in DNA replication / DNA replication origin binding / DNA replication initiation / meiotic cell cycle / DNA helicase activity / mitotic spindle organization / regulation of DNA-templated transcription elongation / helicase activity / mitotic cell cycle / single-stranded DNA binding / DNA replication / DNA helicase / cell division / chromatin binding / ATP hydrolysis activity / ATP binding / nucleus / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
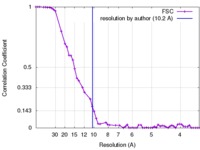
| Method | single particle reconstruction / cryo EM / Resolution: 10.2 Å | |||||||||
 Authors Authors | Renault L / Costa A | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2016 Journal: Nat Commun / Year: 2016Title: Cryo-EM structures of the eukaryotic replicative helicase bound to a translocation substrate. Authors: Ferdos Abid Ali / Ludovic Renault / Julian Gannon / Hailey L Gahlon / Abhay Kotecha / Jin Chuan Zhou / David Rueda / Alessandro Costa /  Abstract: The Cdc45-MCM-GINS (CMG) helicase unwinds DNA during the elongation step of eukaryotic genome duplication and this process depends on the MCM ATPase function. Whether CMG translocation occurs on ...The Cdc45-MCM-GINS (CMG) helicase unwinds DNA during the elongation step of eukaryotic genome duplication and this process depends on the MCM ATPase function. Whether CMG translocation occurs on single- or double-stranded DNA and how ATP hydrolysis drives DNA unwinding remain open questions. Here we use cryo-electron microscopy to describe two subnanometre resolution structures of the CMG helicase trapped on a DNA fork. In the predominant state, the ring-shaped C-terminal ATPase of MCM is compact and contacts single-stranded DNA, via a set of pre-sensor 1 hairpins that spiral around the translocation substrate. In the second state, the ATPase module is relaxed and apparently substrate free, while DNA intimately contacts the downstream amino-terminal tier of the MCM motor ring. These results, supported by single-molecule FRET measurements, lead us to suggest a replication fork unwinding mechanism whereby the N-terminal and AAA+ tiers of the MCM work in concert to translocate on single-stranded DNA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3320.map.gz emd_3320.map.gz | 3.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3320-v30.xml emd-3320-v30.xml emd-3320.xml emd-3320.xml | 20.3 KB 20.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3320_fsc.xml emd_3320_fsc.xml | 5 KB | Display |  FSC data file FSC data file |
| Images |  emd_3320.png emd_3320.png | 1.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3320 http://ftp.pdbj.org/pub/emdb/structures/EMD-3320 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3320 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3320 | HTTPS FTP |
-Validation report
| Summary document |  emd_3320_validation.pdf.gz emd_3320_validation.pdf.gz | 240.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3320_full_validation.pdf.gz emd_3320_full_validation.pdf.gz | 239.9 KB | Display | |
| Data in XML |  emd_3320_validation.xml.gz emd_3320_validation.xml.gz | 8.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3320 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3320 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3320 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3320 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3320.map.gz / Format: CCP4 / Size: 17.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3320.map.gz / Format: CCP4 / Size: 17.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CMG treated with ATP in a tight Mcm5-2 AAA+ configuration | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.77 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : CMG treated with ATP
+Supramolecule #1000: CMG treated with ATP
+Macromolecule #1: Mcm2
+Macromolecule #2: Mcm3
+Macromolecule #3: Mcm4
+Macromolecule #4: Mcm5
+Macromolecule #5: Mcm6
+Macromolecule #6: Mcm7
+Macromolecule #7: Cdc45
+Macromolecule #8: Psf1
+Macromolecule #9: Psf2
+Macromolecule #10: Psf3
+Macromolecule #11: Sld5
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 Details: 25 mM Hepes, 50 mM sodium acetate, 10 mM magnesium acetate, 1 mM DTT, 1mM ATP |
|---|---|
| Grid | Details: 400 mesh quantifoils 1.2/1.3 open holes |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: FEI VITROBOT MARK IV / Method: Blot for 5 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Date | Dec 4, 2014 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON II (4k x 4k) / Digitization - Sampling interval: 14 µm / Number real images: 536 / Average electron dose: 51 e/Å2 / Details: data was recorded as movies of 51 frames / Bits/pixel: 32 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 unidentified baculovirus
unidentified baculovirus