+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
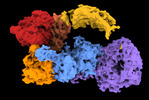
| Title | Structure of the leucine-rich repeat kinase 1 monomer | |||||||||
 Map data Map data | Composite map used for refinement | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  kinase / LRRK / kinase / LRRK /  multi-domain protein / multi-domain protein /  GTPase / GTPase /  TRANSFERASE TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of peptidyl-tyrosine phosphorylation / osteoclast development / positive regulation of intracellular signal transduction /  bone resorption / positive regulation of canonical Wnt signaling pathway / positive regulation of peptidyl-tyrosine phosphorylation / bone resorption / positive regulation of canonical Wnt signaling pathway / positive regulation of peptidyl-tyrosine phosphorylation /  non-specific serine/threonine protein kinase / intracellular signal transduction / non-specific serine/threonine protein kinase / intracellular signal transduction /  phosphorylation / protein serine kinase activity ...negative regulation of peptidyl-tyrosine phosphorylation / osteoclast development / positive regulation of intracellular signal transduction / phosphorylation / protein serine kinase activity ...negative regulation of peptidyl-tyrosine phosphorylation / osteoclast development / positive regulation of intracellular signal transduction /  bone resorption / positive regulation of canonical Wnt signaling pathway / positive regulation of peptidyl-tyrosine phosphorylation / bone resorption / positive regulation of canonical Wnt signaling pathway / positive regulation of peptidyl-tyrosine phosphorylation /  non-specific serine/threonine protein kinase / intracellular signal transduction / non-specific serine/threonine protein kinase / intracellular signal transduction /  phosphorylation / protein serine kinase activity / protein serine/threonine kinase activity / GTP binding / phosphorylation / protein serine kinase activity / protein serine/threonine kinase activity / GTP binding /  mitochondrion / mitochondrion /  ATP binding / identical protein binding / ATP binding / identical protein binding /  metal ion binding / metal ion binding /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.92 Å cryo EM / Resolution: 3.92 Å | |||||||||
 Authors Authors | Metcalfe RD / Martinez Fiesco JA / Zhang P | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structure and regulation of full-length human leucine-rich repeat kinase 1. Authors: Riley D Metcalfe / Juliana A Martinez Fiesco / Luis Bonet-Ponce / Jillian H Kluss / Mark R Cookson / Ping Zhang /  Abstract: The human leucine-rich repeat kinases (LRRKs), LRRK1 and LRRK2 are large and unusually complex multi-domain kinases, which regulate fundamental cellular processes and have been implicated in human ...The human leucine-rich repeat kinases (LRRKs), LRRK1 and LRRK2 are large and unusually complex multi-domain kinases, which regulate fundamental cellular processes and have been implicated in human disease. Structures of LRRK2 have recently been determined, but the structure and molecular mechanisms regulating the activity of the LRRK1 as well as differences in the regulation of LRRK1 and LRRK2 remain unclear. Here, we report a cryo-EM structure of the LRRK1 monomer and a lower-resolution cryo-EM map of the LRRK1 dimer. The monomer structure, in which the kinase is in an inactive conformation, reveals key interdomain interfaces that control kinase activity as we validate experimentally. Both the LRRK1 monomer and dimer are structurally distinct compared to LRRK2. Overall, our results provide structural insights into the activation of the human LRRKs, which advance our understanding of their physiological and pathological roles. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28950.map.gz emd_28950.map.gz | 277.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28950-v30.xml emd-28950-v30.xml emd-28950.xml emd-28950.xml | 13 KB 13 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28950.png emd_28950.png | 99.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28950 http://ftp.pdbj.org/pub/emdb/structures/EMD-28950 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28950 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28950 | HTTPS FTP |
-Related structure data
| Related structure data |  8facMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28950.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28950.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Composite map used for refinement | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.81 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Leucine-rich repeat kinase 1
| Entire | Name: Leucine-rich repeat kinase 1 |
|---|---|
| Components |
|
-Supramolecule #1: Leucine-rich repeat kinase 1
| Supramolecule | Name: Leucine-rich repeat kinase 1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 230 KDa |
-Macromolecule #1: Leucine-rich repeat serine/threonine-protein kinase 1
| Macromolecule | Name: Leucine-rich repeat serine/threonine-protein kinase 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number:  non-specific serine/threonine protein kinase non-specific serine/threonine protein kinase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 229.556359 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MDYKDHDGDY KDHDIDYKDD DDKLGLEVLF QGPMAGMSQR PPSMYWCVGP EESAVCPERA METLNGAGDT GGKPSTRGGD PAARSRRTE GIRAAYRRGD RGGARDLLEE ACDQCASQLE KGQLLSIPAA YGDLEMVRYL LSKRLVELPT EPTDDNPAVV A AYFGHTAV ...String: MDYKDHDGDY KDHDIDYKDD DDKLGLEVLF QGPMAGMSQR PPSMYWCVGP EESAVCPERA METLNGAGDT GGKPSTRGGD PAARSRRTE GIRAAYRRGD RGGARDLLEE ACDQCASQLE KGQLLSIPAA YGDLEMVRYL LSKRLVELPT EPTDDNPAVV A AYFGHTAV VQELLESLPG PCSPQRLLNW MLALACQRGH LGVVKLLVLT HGADPESYAV RKNEFPVIVR LPLYAAIKSG NE DIAIFLL RHGAYFCSYI LLDSPDPSKH LLRKYFIEAS PLPSSYPGKT ALRVKWSHLR LPWVDLDWLI DISCQITELD LSA NCLATL PSVIPWGLIN LRKLNLSDNH LGELPGVQSS DEIICSRLLE IDISSNKLSH LPPGFLHLSK LQKLTASKNC LEKL FEEEN ATNWIGLRKL QELDISDNKL TELPALFLHS FKSLNSLNVS RNNLKVFPDP WACPLKCCKA SRNALECLPD KMAVF WKNH LKDVDFSENA LKEVPLGLFQ LDALMFLRLQ GNQLAALPPQ EKWTCRQLKT LDLSRNQLGK NEDGLKTKRI AFFTTR GRQ RSGTEAASVL EFPAFLSESL EVLCLNDNHL DTVPPSVCLL KSLSELYLGN NPGLRELPPE LGQLGNLWQL DTEDLTI SN VPAEIQKEGP KAMLSYLRAQ LRKAEKCKLM KMIIVGPPRQ GKSTLLEILQ TGRAPQVVHG EATIRTTKWE LQRPAGSR A KVESVEFNVW DIGGPASMAT VNQCFFTDKA LYVVVWNLAL GEEAVANLQF WLLNIEAKAP NAVVLVVGTH LDLIEAKFR VERIATLRAY VLALCRSPSG SRATGFPDIT FKHLHEISCK SLEGQEGLRQ LIFHVTCSMK DVGSTIGCQR LAGRLIPRSY LSLQEAVLA EQQRRSRDDD VQYLTDRQLE QLVEQTPDND IKDYEDLQSA ISFLIETGTL LHFPDTSHGL RNLYFLDPIW L SECLQRIF NIKGSRSVAK NGVIRAEDLR MLLVGTGFTQ QTEEQYFQFL AKFEIALPVA NDSYLLPHLL PSKPGLDTHG MR HPTANTI QRVFKMSFVP VGFWQRFIAR MLISLAEMDL QLFENKKNTK SRNRKVTIYS FTGNQRNRCS TFRVKRNQTI YWQ EGLLVT FDGGYLSVES SDVNWKKKKS GGMKIVCQSE VRDFSAMAFI TDHVNSLIDQ WFPALTATES DGTPLMEQYV PCPV CETAW AQHTDPSEKS EDVQYFDMED CVLTAIERDF ISCPRHPDLP VPLQELVPEL FMTDFPARLF LENSKLEHSE DEGSV LGQG GSGTVIYRAR YQGQPVAVKR FHIKKFKNFA NVPADTMLRH LRATDAMKNF SEFRQEASML HALQHPCIVA LIGISI HPL CFALELAPLS SLNTVLSENA RDSSFIPLGH MLTQKIAYQI ASGLAYLHKK NIIFCDLKSD NILVWSLDVK EHINIKL SD YGISRQSFHE GALGVEGTPG YQAPEIRPRI VYDEKVDMFS YGMVLYELLS GQRPALGHHQ LQIAKKLSKG IRPVLGQP E EVQFRRLQAL MMECWDTKPE KRPLALSVVS QMKDPTFATF MYELCCGKQT AFFSSQGQEY TVVFWDGKEE SRNYTVVNT EKGLMEVQRM CCPGMKVSCQ LQVQRSLWTA TEDQKIYIYT LKGMCPLNTP QQALDTPAVV TCFLAVPVIK KNSYLVLAGL ADGLVAVFP VVRGTPKDSC SYLCSHTANR SKFSIADEDA RQNPYPVKAM EVVNSGSEVW YSNGPGLLVI DCASLEICRR L EPYMAPSM VTSVVCSSEG RGEEVVWCLD DKANSLVMYH STTYQLCARY FCGVPSPLRD MFPVRPLDTE PPAASHTANP KV PEGDSIA DVSIMYSEEL GTQILIHQES LTDYCSMSSY SSSPPRQAAR SPSSLPSSPA SSSSVPFSTD CEDSDMLHTP GAA SDRSEH DLTPMDGETF SQHLQAVKIL AVRDLIWVPR RGGDVIVIGL EKDSGAQRGR VIAVLKAREL TPHGVLVDAA VVAK DTVVC TFENENTEWC LAVWRGWGAR EFDIFYQSYE ELGRLEACTR KRR UniProtKB: Leucine-rich repeat serine/threonine-protein kinase 1 |
-Macromolecule #2: GUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 1 / Formula: GDP |
|---|---|
| Molecular weight | Theoretical: 443.201 Da |
| Chemical component information |  ChemComp-GDP: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 8.3 |
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA EM GP |
| Details | Purified LRRK1 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 7 / Number real images: 18074 / Average exposure time: 2.5 sec. / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 3553719 Details: Particles picked using a Topaz model trained on an initial LRRK1 dataset. |
|---|---|
| Startup model | Type of model: NONE / Details: Ab-initio model generated using Cryosparc. |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.92 Å / Resolution method: FSC 0.143 CUT-OFF Details: Resolution given for map is the resolution of the global refinement map (D_1000269296). The resolution for the local refinement map was 3.94 (D_1000269298). Number images used: 183273 |
 Movie
Movie Controller
Controller