[English] 日本語
 Yorodumi
Yorodumi- EMDB-27641: The structure of the interleukin 11 signalling complex, truncated... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of the interleukin 11 signalling complex, truncated gp130 | |||||||||
 Map data Map data | The structure of the interleukin 11 signalling complex, truncated gp130 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  complex / IL-6 family cytokine / gp130 / glycoprotein 130 / complex / IL-6 family cytokine / gp130 / glycoprotein 130 /  glycoprotein / glycoprotein /  CYTOKINE CYTOKINE | |||||||||
| Function / homology |  Function and homology information Function and homology information interleukin-11 receptor binding / interleukin-11 receptor binding /  interleukin-27 receptor activity / oncostatin-M receptor complex / oncostatin-M-mediated signaling pathway / interleukin-27 receptor activity / oncostatin-M receptor complex / oncostatin-M-mediated signaling pathway /  ciliary neurotrophic factor receptor activity / ciliary neurotrophic factor receptor activity /  ciliary neurotrophic factor receptor binding / negative regulation of interleukin-6-mediated signaling pathway / leukemia inhibitory factor signaling pathway / ciliary neurotrophic factor receptor binding / negative regulation of interleukin-6-mediated signaling pathway / leukemia inhibitory factor signaling pathway /  interleukin-11 receptor activity / interleukin-11 receptor activity /  interleukin-11 binding ... interleukin-11 binding ... interleukin-11 receptor binding / interleukin-11 receptor binding /  interleukin-27 receptor activity / oncostatin-M receptor complex / oncostatin-M-mediated signaling pathway / interleukin-27 receptor activity / oncostatin-M receptor complex / oncostatin-M-mediated signaling pathway /  ciliary neurotrophic factor receptor activity / ciliary neurotrophic factor receptor activity /  ciliary neurotrophic factor receptor binding / negative regulation of interleukin-6-mediated signaling pathway / leukemia inhibitory factor signaling pathway / ciliary neurotrophic factor receptor binding / negative regulation of interleukin-6-mediated signaling pathway / leukemia inhibitory factor signaling pathway /  interleukin-11 receptor activity / interleukin-11 receptor activity /  interleukin-11 binding / megakaryocyte differentiation / interleukin-11 binding / megakaryocyte differentiation /  ciliary neurotrophic factor receptor complex / interleukin-27-mediated signaling pathway / ciliary neurotrophic factor-mediated signaling pathway / ciliary neurotrophic factor receptor complex / interleukin-27-mediated signaling pathway / ciliary neurotrophic factor-mediated signaling pathway /  interleukin-6 receptor complex / head development / negative regulation of hormone secretion / interleukin-11-mediated signaling pathway / T-helper 17 cell lineage commitment / positive regulation of adaptive immune response / interleukin-6 receptor complex / head development / negative regulation of hormone secretion / interleukin-11-mediated signaling pathway / T-helper 17 cell lineage commitment / positive regulation of adaptive immune response /  developmental process / positive regulation of acute inflammatory response / positive regulation of astrocyte differentiation / intestinal epithelial cell development / positive regulation of platelet aggregation / Interleukin-35 Signalling / Interleukin-27 signaling / IL-6-type cytokine receptor ligand interactions / developmental process / positive regulation of acute inflammatory response / positive regulation of astrocyte differentiation / intestinal epithelial cell development / positive regulation of platelet aggregation / Interleukin-35 Signalling / Interleukin-27 signaling / IL-6-type cytokine receptor ligand interactions /  cytokine receptor activity / glycogen metabolic process / Interleukin-6 signaling / interleukin-6-mediated signaling pathway / positive regulation of Notch signaling pathway / protein tyrosine kinase activator activity / cytokine receptor activity / glycogen metabolic process / Interleukin-6 signaling / interleukin-6-mediated signaling pathway / positive regulation of Notch signaling pathway / protein tyrosine kinase activator activity /  cytokine binding / positive regulation of cardiac muscle hypertrophy / fat cell differentiation / MAPK3 (ERK1) activation / cytokine binding / positive regulation of cardiac muscle hypertrophy / fat cell differentiation / MAPK3 (ERK1) activation /  growth factor binding / MAPK1 (ERK2) activation / positive regulation of vascular endothelial growth factor production / positive regulation of osteoblast differentiation / growth factor binding / MAPK1 (ERK2) activation / positive regulation of vascular endothelial growth factor production / positive regulation of osteoblast differentiation /  coreceptor activity / positive regulation of tyrosine phosphorylation of STAT protein / positive regulation of T cell proliferation / B cell differentiation / response to cytokine / coreceptor activity / positive regulation of tyrosine phosphorylation of STAT protein / positive regulation of T cell proliferation / B cell differentiation / response to cytokine /  cytokine activity / cytokine activity /  growth factor activity / cytokine-mediated signaling pathway / positive regulation of peptidyl-tyrosine phosphorylation / transmembrane signaling receptor activity / positive regulation of peptidyl-serine phosphorylation / growth factor activity / cytokine-mediated signaling pathway / positive regulation of peptidyl-tyrosine phosphorylation / transmembrane signaling receptor activity / positive regulation of peptidyl-serine phosphorylation /  scaffold protein binding / cell population proliferation / negative regulation of neuron apoptotic process / positive regulation of MAPK cascade / scaffold protein binding / cell population proliferation / negative regulation of neuron apoptotic process / positive regulation of MAPK cascade /  receptor complex / receptor complex /  membrane raft / external side of plasma membrane / neuronal cell body / membrane raft / external side of plasma membrane / neuronal cell body /  dendrite / positive regulation of cell population proliferation / negative regulation of apoptotic process / positive regulation of transcription by RNA polymerase II / dendrite / positive regulation of cell population proliferation / negative regulation of apoptotic process / positive regulation of transcription by RNA polymerase II /  extracellular space / extracellular exosome / extracellular region / extracellular space / extracellular exosome / extracellular region /  membrane / identical protein binding / membrane / identical protein binding /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
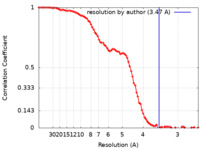
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.47 Å cryo EM / Resolution: 3.47 Å | |||||||||
 Authors Authors | Metcalfe RD / Hanssen E / Griffin MDW | |||||||||
| Funding support |  Australia, 1 items Australia, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structures of the interleukin 11 signalling complex reveal gp130 dynamics and the inhibitory mechanism of a cytokine variant. Authors: Riley D Metcalfe / Eric Hanssen / Ka Yee Fung / Kaheina Aizel / Clara C Kosasih / Courtney O Zlatic / Larissa Doughty / Craig J Morton / Andrew P Leis / Michael W Parker / Paul R Gooley / ...Authors: Riley D Metcalfe / Eric Hanssen / Ka Yee Fung / Kaheina Aizel / Clara C Kosasih / Courtney O Zlatic / Larissa Doughty / Craig J Morton / Andrew P Leis / Michael W Parker / Paul R Gooley / Tracy L Putoczki / Michael D W Griffin /   Abstract: Interleukin (IL-)11, an IL-6 family cytokine, has pivotal roles in autoimmune diseases, fibrotic complications, and solid cancers. Despite intense therapeutic targeting efforts, structural ...Interleukin (IL-)11, an IL-6 family cytokine, has pivotal roles in autoimmune diseases, fibrotic complications, and solid cancers. Despite intense therapeutic targeting efforts, structural understanding of IL-11 signalling and mechanistic insights into current inhibitors are lacking. Here we present cryo-EM and crystal structures of the human IL-11 signalling complex, including the complex containing the complete extracellular domains of the shared IL-6 family β-receptor, gp130. We show that complex formation requires conformational reorganisation of IL-11 and that the membrane-proximal domains of gp130 are dynamic. We demonstrate that the cytokine mutant, IL-11 Mutein, competitively inhibits signalling in human cell lines. Structural shifts in IL-11 Mutein underlie inhibition by altering cytokine binding interactions at all three receptor-engaging sites and abrogating the final gp130 binding step. Our results reveal the structural basis of IL-11 signalling, define the molecular mechanisms of an inhibitor, and advance understanding of gp130-containing receptor complexes, with potential applications in therapeutic development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27641.map.gz emd_27641.map.gz | 48.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27641-v30.xml emd-27641-v30.xml emd-27641.xml emd-27641.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27641_fsc.xml emd_27641_fsc.xml | 10.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_27641.png emd_27641.png | 115.5 KB | ||
| Filedesc metadata |  emd-27641.cif.gz emd-27641.cif.gz | 6.2 KB | ||
| Others |  emd_27641_half_map_1.map.gz emd_27641_half_map_1.map.gz emd_27641_half_map_2.map.gz emd_27641_half_map_2.map.gz | 38.8 MB 38.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27641 http://ftp.pdbj.org/pub/emdb/structures/EMD-27641 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27641 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27641 | HTTPS FTP |
-Related structure data
| Related structure data |  8dpsMC  8dptC  8dpuC  8dpvC  8dpwC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27641.map.gz / Format: CCP4 / Size: 51.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27641.map.gz / Format: CCP4 / Size: 51.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The structure of the interleukin 11 signalling complex, truncated gp130 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: The structure of the interleukin 11 signalling complex,...
| File | emd_27641_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The structure of the interleukin 11 signalling complex, truncated gp130 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: The structure of the interleukin 11 signalling complex,...
| File | emd_27641_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The structure of the interleukin 11 signalling complex, truncated gp130 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Interleukin 11 signalling complex
| Entire | Name: Interleukin 11 signalling complex |
|---|---|
| Components |
|
-Supramolecule #1: Interleukin 11 signalling complex
| Supramolecule | Name: Interleukin 11 signalling complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 / Details: 2:2:2 hexameric complex |
|---|---|
| Molecular weight | Theoretical: 182 KDa |
-Supramolecule #2: gp130
| Supramolecule | Name: gp130 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Interleukin-11 receptor subunit alpha
| Supramolecule | Name: Interleukin-11 receptor subunit alpha / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #4: Interleukin-11
| Supramolecule | Name: Interleukin-11 / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Interleukin-6 receptor subunit beta
| Macromolecule | Name: Interleukin-6 receptor subunit beta / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 34.563961 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: GELLDPCGYI SPESPVVQLH SNFTAVCVLK EKCMDYFHVN ANYIVWKTNH FTIPKEQYTI INRTASSVTF TDIASLNIQL TCNILTFGQ LEQNVYGITI ISGLPPEKPK NLSCIVNEGK KMRCEWDGGR ETHLETNFTL KSEWATHKFA DCKAKRDTPT S CTVDYSTV ...String: GELLDPCGYI SPESPVVQLH SNFTAVCVLK EKCMDYFHVN ANYIVWKTNH FTIPKEQYTI INRTASSVTF TDIASLNIQL TCNILTFGQ LEQNVYGITI ISGLPPEKPK NLSCIVNEGK KMRCEWDGGR ETHLETNFTL KSEWATHKFA DCKAKRDTPT S CTVDYSTV YFVNIEVWVE AENALGKVTS DHINFDPVYK VKPNPPHNLS VINSEELSSI LKLTWTNPSI KSVIILKYNI QY RTKDAST WSQIPPEDTA STRSSFTVQD LKPFTEYVFR IRCMKEDGKG YWSDWSEEAS GITYED UniProtKB:  Interleukin-6 receptor subunit beta Interleukin-6 receptor subunit beta |
-Macromolecule #2: Interleukin-11
| Macromolecule | Name: Interleukin-11 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.273289 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: GSPDPRAELD STVLLTRSLL ADTRQLAAQL RDKFPADGDH NLDSLPTLAM SAGALGALQL PGVLTRLRAD LLSYLRHVQW LRRAGGSSL KTLEPELGTL QARLDRLLRR LQLLMSRLAL PQPPPDPPAP PLAPPSSAWG GIRAAHAILG GLHLTLDWAV R GLLLLKTR L UniProtKB:  Interleukin-11 Interleukin-11 |
-Macromolecule #3: Interleukin-11 receptor subunit alpha
| Macromolecule | Name: Interleukin-11 receptor subunit alpha / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32.208002 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: GSSPCPQAWG PPGVQYGQPG RSVKLCCPGV TAGDPVSWFR DGEPKLLQGP DSGLGHELVL AQADSTDEGT YICQTLDGAL GGTVTLQLG YPPARPVVSC QAADYENFSC TWSPSQISGL PTRYLTSYRK KTVLGADSQR RSPSTGPWPC PQDPLGAARC V VHGAEFWS ...String: GSSPCPQAWG PPGVQYGQPG RSVKLCCPGV TAGDPVSWFR DGEPKLLQGP DSGLGHELVL AQADSTDEGT YICQTLDGAL GGTVTLQLG YPPARPVVSC QAADYENFSC TWSPSQISGL PTRYLTSYRK KTVLGADSQR RSPSTGPWPC PQDPLGAARC V VHGAEFWS QYRINVTEVN PLGASTRLLD VSLQSILRPD PPQGLRVESV PGYPRRLRAS WTYPASWPSQ PHFLLKFRLQ YR PAQHPAW STVEPAGLEE VITDAVAGLP HAVRVSARDF LDAGTWSTWS PEAWGTPSTG T UniProtKB:  Interleukin-11 receptor subunit alpha Interleukin-11 receptor subunit alpha |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 4 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8.5 Component:
Details: TBS pH 85 | |||||||||
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 52.0 e/Å2 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z
Z Y
Y X
X