+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2635 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron microscopy of human transcriptional Mediator | |||||||||
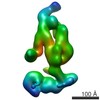
 Map data Map data | Negative-stained reconstruction of human transcriptional Mediator | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | human / transcription / Mediator / Med26 / RNAPII / MED | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 30.0 Å | |||||||||
 Authors Authors | Tsai KT / Tomomori-Sato C / Sato S / Conaway RC / Conaway JW / Asturias FJ | |||||||||
 Citation Citation |  Journal: Cell / Year: 2014 Journal: Cell / Year: 2014Title: Subunit architecture and functional modular rearrangements of the transcriptional mediator complex. Authors: Kuang-Lei Tsai / Chieri Tomomori-Sato / Shigeo Sato / Ronald C Conaway / Joan W Conaway / Francisco J Asturias /  Abstract: The multisubunit Mediator, comprising ∼30 distinct proteins, plays an essential role in gene expression regulation by acting as a bridge between DNA-binding transcription factors and the RNA ...The multisubunit Mediator, comprising ∼30 distinct proteins, plays an essential role in gene expression regulation by acting as a bridge between DNA-binding transcription factors and the RNA polymerase II (RNAPII) transcription machinery. Efforts to uncover the Mediator mechanism have been hindered by a poor understanding of its structure, subunit organization, and conformational rearrangements. By overcoming biochemical and image analysis hurdles, we obtained accurate EM structures of yeast and human Mediators. Subunit localization experiments, docking of partial X-ray structures, and biochemical analyses resulted in comprehensive mapping of yeast Mediator subunits and a complete reinterpretation of our previous Mediator organization model. Large-scale Mediator rearrangements depend on changes at the interfaces between previously described Mediator modules, which appear to be facilitated by factors conducive to transcription initiation. Conservation across eukaryotes of Mediator structure, subunit organization, and RNA polymerase II interaction suggest conservation of fundamental aspects of the Mediator mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2635.map.gz emd_2635.map.gz | 7.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2635-v30.xml emd-2635-v30.xml emd-2635.xml emd-2635.xml | 7.3 KB 7.3 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2635_hMED.jpg EMD-2635_hMED.jpg | 63.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2635 http://ftp.pdbj.org/pub/emdb/structures/EMD-2635 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2635 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2635 | HTTPS FTP |
-Validation report
| Summary document |  emd_2635_validation.pdf.gz emd_2635_validation.pdf.gz | 210.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2635_full_validation.pdf.gz emd_2635_full_validation.pdf.gz | 209.5 KB | Display | |
| Data in XML |  emd_2635_validation.xml.gz emd_2635_validation.xml.gz | 4.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2635 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2635 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2635 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2635 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2635.map.gz / Format: CCP4 / Size: 7.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2635.map.gz / Format: CCP4 / Size: 7.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Negative-stained reconstruction of human transcriptional Mediator | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human transcriptional Mediator
| Entire | Name: human transcriptional Mediator |
|---|---|
| Components |
|
-Supramolecule #1000: human transcriptional Mediator
| Supramolecule | Name: human transcriptional Mediator / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Macromolecule #1: human transcriptional Mediator
| Macromolecule | Name: human transcriptional Mediator / type: protein_or_peptide / ID: 1 / Name.synonym: hMED / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Strain: HeLa-S3 / synonym: Human Homo sapiens (human) / Strain: HeLa-S3 / synonym: Human |
| Molecular weight | Theoretical: 1 MDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Staining | Type: NEGATIVE Details: Grids with adsorbed protein floated on 0.75% w/v uranyl formate for 30 seconds |
|---|---|
| Grid | Details: 400 mesh gold grid with thin carbon support |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Date | Jun 10, 2013 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F415 (4k x 4k) |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 52000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 30.0 Å / Resolution method: OTHER / Software - Name: EMAN2/SPARX / Number images used: 2025 |
|---|
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera