[English] 日本語
 Yorodumi
Yorodumi- EMDB-1896: EM map of the specific p53-DNA complex at 21 angstroms resolution -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1896 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | EM map of the specific p53-DNA complex at 21 angstroms resolution | |||||||||
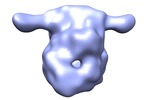
 Map data Map data | 3D EM map of the specific p53-DNA complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cryo electron microscopy / p53 / single particle analysis / transcription factor | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 21.0 Å | |||||||||
 Authors Authors | Aramayo R / Sherman MB / Brownless K / Lurz R / Okorokov AL / Orlova EV | |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2011 Journal: Nucleic Acids Res / Year: 2011Title: Quaternary structure of the specific p53-DNA complex reveals the mechanism of p53 mutant dominance. Authors: Ricardo Aramayo / Michael B Sherman / Kathryne Brownless / Rudi Lurz / Andrei L Okorokov / Elena V Orlova /  Abstract: The p53 tumour suppressor is a transcriptional activator that controls cell fate in response to various stresses. p53 can initiate cell cycle arrest, senescence and/or apoptosis via transactivation ...The p53 tumour suppressor is a transcriptional activator that controls cell fate in response to various stresses. p53 can initiate cell cycle arrest, senescence and/or apoptosis via transactivation of p53 target genes, thus preventing cancer onset. Mutations that impair p53 usually occur in the core domain and negate the p53 sequence-specific DNA binding. Moreover, these mutations exhibit a dominant negative effect on the remaining wild-type p53. Here, we report the cryo electron microscopy structure of the full-length p53 tetramer bound to a DNA-encoding transcription factor response element (RE) at a resolution of 21 A. While two core domains from both dimers of the p53 tetramer interact with DNA within the complex, the other two core domains remain available for binding another DNA site. This finding helps to explain the dominant negative effect of p53 mutants based on the fact that p53 dimers are formed co-translationally before the whole tetramer assembles; therefore, a single mutant dimer would prevent the p53 tetramer from binding DNA. The structure indicates that the Achilles' heel of p53 is in its dimer-of-dimers organization, thus the tetramer activity can be negated by mutation in only one allele followed by tumourigenesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1896.map.gz emd_1896.map.gz | 395.9 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1896-v30.xml emd-1896-v30.xml emd-1896.xml emd-1896.xml | 8.6 KB 8.6 KB | Display Display |  EMDB header EMDB header |
| Images |  em-1896.png em-1896.png | 248.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1896 http://ftp.pdbj.org/pub/emdb/structures/EMD-1896 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1896 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1896 | HTTPS FTP |
-Validation report
| Summary document |  emd_1896_validation.pdf.gz emd_1896_validation.pdf.gz | 190.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1896_full_validation.pdf.gz emd_1896_full_validation.pdf.gz | 190 KB | Display | |
| Data in XML |  emd_1896_validation.xml.gz emd_1896_validation.xml.gz | 5.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1896 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1896 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1896 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1896 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1896.map.gz / Format: CCP4 / Size: 2.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1896.map.gz / Format: CCP4 / Size: 2.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D EM map of the specific p53-DNA complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.48 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Murine p53 tetramer complexed with specific DNA construct
| Entire | Name: Murine p53 tetramer complexed with specific DNA construct |
|---|---|
| Components |
|
-Supramolecule #1000: Murine p53 tetramer complexed with specific DNA construct
| Supramolecule | Name: Murine p53 tetramer complexed with specific DNA construct type: sample / ID: 1000 / Number unique components: 2 |
|---|---|
| Molecular weight | Experimental: 240 KDa / Theoretical: 240 KDa |
-Macromolecule #1: p53
| Macromolecule | Name: p53 / type: protein_or_peptide / ID: 1 / Name.synonym: p53 Details: Murine p53 tetramer bound to a specific DNA construct Number of copies: 4 / Oligomeric state: Tetramer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 240 KDa / Theoretical: 240 KDa |
| Recombinant expression | Organism:  unidentified baculovirus unidentified baculovirus |
-Macromolecule #2: DNA
| Macromolecule | Name: DNA / type: dna / ID: 2 / Name.synonym: DNA / Classification: DNA / Structure: DOUBLE HELIX / Synthetic?: No |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 / Details: 100mM NaCl, 25mM Tris-HCl, 1mM DTT |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 80 K / Instrument: OTHER / Method: Blot few seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2200FS |
|---|---|
| Temperature | Average: 90 K |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Average electron dose: 10 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 5.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: Phase flipping |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 21.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: IMAGIC, EMAN |
 Movie
Movie Controller
Controller