[English] 日本語
 Yorodumi
Yorodumi- EMDB-15113: Human mature large subunit of the ribosome with eIF6 and homoharr... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human mature large subunit of the ribosome with eIF6 and homoharringtonine bound | |||||||||
 Map data Map data | Postprocessed map. Masked, and sharpened at -40 A2. Must be upsampled 768x768x768 (0.55267 pixel size) for best visualisation and alignment with supplied structure. | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationlamin filament / regulation of fatty acid biosynthetic process / regulation of megakaryocyte differentiation / miRNA-mediated post-transcriptional gene silencing / embryonic brain development / eukaryotic 80S initiation complex / negative regulation of protein neddylation / miRNA-mediated gene silencing by inhibition of translation /  translation at presynapse / axial mesoderm development ...lamin filament / regulation of fatty acid biosynthetic process / regulation of megakaryocyte differentiation / miRNA-mediated post-transcriptional gene silencing / embryonic brain development / eukaryotic 80S initiation complex / negative regulation of protein neddylation / miRNA-mediated gene silencing by inhibition of translation / translation at presynapse / axial mesoderm development ...lamin filament / regulation of fatty acid biosynthetic process / regulation of megakaryocyte differentiation / miRNA-mediated post-transcriptional gene silencing / embryonic brain development / eukaryotic 80S initiation complex / negative regulation of protein neddylation / miRNA-mediated gene silencing by inhibition of translation /  translation at presynapse / axial mesoderm development / regulation of G1 to G0 transition / negative regulation of formation of translation preinitiation complex / translation at presynapse / axial mesoderm development / regulation of G1 to G0 transition / negative regulation of formation of translation preinitiation complex /  ribosomal protein import into nucleus / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / regulation of translation involved in cellular response to UV / exit from mitosis / protein-DNA complex disassembly / 90S preribosome assembly / positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator / optic nerve development / TORC2 complex binding / ribosomal protein import into nucleus / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / regulation of translation involved in cellular response to UV / exit from mitosis / protein-DNA complex disassembly / 90S preribosome assembly / positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator / optic nerve development / TORC2 complex binding /  GAIT complex / G1 to G0 transition / retinal ganglion cell axon guidance / middle ear morphogenesis / cytoplasmic side of rough endoplasmic reticulum membrane / A band / regulation of reactive oxygen species metabolic process / positive regulation of signal transduction by p53 class mediator / alpha-beta T cell differentiation / ubiquitin ligase inhibitor activity / regulation of glycolytic process / negative regulation of ubiquitin protein ligase activity / response to aldosterone / maturation of 5.8S rRNA / homeostatic process / lung morphogenesis / macrophage chemotaxis / GAIT complex / G1 to G0 transition / retinal ganglion cell axon guidance / middle ear morphogenesis / cytoplasmic side of rough endoplasmic reticulum membrane / A band / regulation of reactive oxygen species metabolic process / positive regulation of signal transduction by p53 class mediator / alpha-beta T cell differentiation / ubiquitin ligase inhibitor activity / regulation of glycolytic process / negative regulation of ubiquitin protein ligase activity / response to aldosterone / maturation of 5.8S rRNA / homeostatic process / lung morphogenesis / macrophage chemotaxis /  ribosomal subunit export from nucleus / ribosomal subunit export from nucleus /  Protein hydroxylation / Peptide chain elongation / Protein hydroxylation / Peptide chain elongation /  ribosomal large subunit binding / Selenocysteine synthesis / protein-RNA complex assembly / Formation of a pool of free 40S subunits / Eukaryotic Translation Termination / blastocyst development / preribosome, large subunit precursor / Response of EIF2AK4 (GCN2) to amino acid deficiency / translation regulator activity / SRP-dependent cotranslational protein targeting to membrane / Viral mRNA Translation / protein localization to nucleus / cellular response to actinomycin D / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / negative regulation of proteasomal ubiquitin-dependent protein catabolic process / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression / ribosomal large subunit binding / Selenocysteine synthesis / protein-RNA complex assembly / Formation of a pool of free 40S subunits / Eukaryotic Translation Termination / blastocyst development / preribosome, large subunit precursor / Response of EIF2AK4 (GCN2) to amino acid deficiency / translation regulator activity / SRP-dependent cotranslational protein targeting to membrane / Viral mRNA Translation / protein localization to nucleus / cellular response to actinomycin D / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / negative regulation of proteasomal ubiquitin-dependent protein catabolic process / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression /  protein targeting / Major pathway of rRNA processing in the nucleolus and cytosol / protein targeting / Major pathway of rRNA processing in the nucleolus and cytosol /  rough endoplasmic reticulum / MDM2/MDM4 family protein binding / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of LSU-rRNA / negative regulation of ubiquitin-dependent protein catabolic process / rough endoplasmic reticulum / MDM2/MDM4 family protein binding / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of LSU-rRNA / negative regulation of ubiquitin-dependent protein catabolic process /  ribosomal large subunit biogenesis / ribosomal large subunit biogenesis /  embryo implantation / cellular response to interleukin-4 / embryo implantation / cellular response to interleukin-4 /  translation initiation factor activity / translation initiation factor activity /  ossification / assembly of large subunit precursor of preribosome / cytosolic ribosome / cytosolic ribosome assembly / positive regulation of translation / ossification / assembly of large subunit precursor of preribosome / cytosolic ribosome / cytosolic ribosome assembly / positive regulation of translation /  innate immune response in mucosa / regulation of signal transduction by p53 class mediator / mRNA 3'-UTR binding / innate immune response in mucosa / regulation of signal transduction by p53 class mediator / mRNA 3'-UTR binding /  skeletal system development / sensory perception of sound / response to insulin / cellular response to gamma radiation / skeletal system development / sensory perception of sound / response to insulin / cellular response to gamma radiation /  bone development / mRNA 5'-UTR binding / bone development / mRNA 5'-UTR binding /  ribosomal large subunit assembly / ribosomal large subunit assembly /  transcription coactivator binding / cytoplasmic ribonucleoprotein granule / rRNA processing / Regulation of expression of SLITs and ROBOs / cellular response to type II interferon / cellular response to UV / antimicrobial humoral immune response mediated by antimicrobial peptide / large ribosomal subunit rRNA binding / transcription coactivator binding / cytoplasmic ribonucleoprotein granule / rRNA processing / Regulation of expression of SLITs and ROBOs / cellular response to type II interferon / cellular response to UV / antimicrobial humoral immune response mediated by antimicrobial peptide / large ribosomal subunit rRNA binding /  ribosome binding / large ribosomal subunit / presynapse / ribosome binding / large ribosomal subunit / presynapse /  regulation of translation / retina development in camera-type eye / regulation of translation / retina development in camera-type eye /  cell body cell bodySimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) / Homo sapiens (human) /   human (human) human (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 1.67 Å cryo EM / Resolution: 1.67 Å | |||||||||
 Authors Authors | Faille A / Warren AJ / Dent KC | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: bioRxiv / Year: 2023 Journal: bioRxiv / Year: 2023Title: The chemical landscape of the human ribosome at 1.67 Å resolution. Authors: Alexandre Faille / Kyle C Dent / Simone Pellegrino / Pekka Jaako / Alan J Warren Abstract: The ability of ribosomes to translate the genetic code into protein requires a finely tuned ion and solvent ecosystem. However, the lack of high-resolution structures has precluded accurate ...The ability of ribosomes to translate the genetic code into protein requires a finely tuned ion and solvent ecosystem. However, the lack of high-resolution structures has precluded accurate positioning of all the functional elements of the ribosome and limited our understanding of the specific role of ribosomal RNA chemical modifications in modulating ribosome function in health and disease. Here, using a new sample preparation methodology based on functionalised pristine graphene-coated grids, we solve the cryo-EM structure of the human large ribosomal subunit to a resolution of 1.67 Å. The accurate assignment of water molecules, magnesium and potassium ions in our model highlights the fundamental biological role of ribosomal RNA methylation in harnessing unconventional carbon-oxygen hydrogen bonds to establish chemical interactions with the environment and fine-tune the functional interplay with tRNA. In addition, the structures of three translational inhibitors bound to the human large ribosomal subunit at better than 2 Å resolution provide mechanistic insights into how three key druggable pockets of the ribosome are targeted and illustrate the potential of this methodology to accelerate high-throughput structure-based design of anti-cancer therapeutics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15113.map.gz emd_15113.map.gz | 58.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15113-v30.xml emd-15113-v30.xml emd-15113.xml emd-15113.xml | 68 KB 68 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15113_fsc.xml emd_15113_fsc.xml | 17.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_15113.png emd_15113.png | 81 KB | ||
| Masks |  emd_15113_msk_1.map emd_15113_msk_1.map | 512 MB |  Mask map Mask map | |
| Others |  emd_15113_additional_1.map.gz emd_15113_additional_1.map.gz emd_15113_half_map_1.map.gz emd_15113_half_map_1.map.gz emd_15113_half_map_2.map.gz emd_15113_half_map_2.map.gz | 405.2 MB 409.1 MB 409.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15113 http://ftp.pdbj.org/pub/emdb/structures/EMD-15113 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15113 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15113 | HTTPS FTP |
-Related structure data
| Related structure data |  8a3dMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15113.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15113.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed map. Masked, and sharpened at -40 A2. Must be upsampled 768x768x768 (0.55267 pixel size) for best visualisation and alignment with supplied structure. | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.829 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15113_msk_1.map emd_15113_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
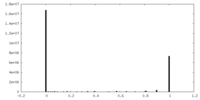
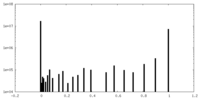
| Density Histograms |
-Additional map: Final refinement map.
| File | emd_15113_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final refinement map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
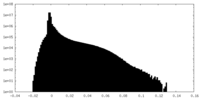
| Density Histograms |
-Half map: Final refinement half-map 1.
| File | emd_15113_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final refinement half-map 1. | ||||||||||||
| Projections & Slices |
| ||||||||||||
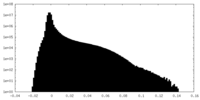
| Density Histograms |
-Half map: Final refinement half-map 2.
| File | emd_15113_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final refinement half-map 2. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Human mature large subunit of the ribosome with eIF6 bound
+Supramolecule #1: Human mature large subunit of the ribosome with eIF6 bound
+Macromolecule #1: 28S ribosomal RNA
+Macromolecule #2: 5S ribosomal RNA
+Macromolecule #3: 5.8S ribosomal RNA
+Macromolecule #4: 60S ribosomal protein L8
+Macromolecule #5: 60S ribosomal protein L3
+Macromolecule #6: 60S ribosomal protein L4
+Macromolecule #7: 60S ribosomal protein L5
+Macromolecule #8: 60S ribosomal protein L6
+Macromolecule #9: 60S ribosomal protein L7
+Macromolecule #10: 60S ribosomal protein L7a
+Macromolecule #11: 60S ribosomal protein L9
+Macromolecule #12: 60S ribosomal protein L10
+Macromolecule #13: 60S ribosomal protein L11
+Macromolecule #14: 60S ribosomal protein L13
+Macromolecule #15: 60S ribosomal protein L14
+Macromolecule #16: 60S ribosomal protein L15
+Macromolecule #17: 60S ribosomal protein L13a
+Macromolecule #18: 60S ribosomal protein L17
+Macromolecule #19: 60S ribosomal protein L18
+Macromolecule #20: 60S ribosomal protein L19
+Macromolecule #21: 60S ribosomal protein L18a
+Macromolecule #22: 60S ribosomal protein L21
+Macromolecule #23: 60S ribosomal protein L22
+Macromolecule #24: 60S ribosomal protein L23
+Macromolecule #25: 60S ribosomal protein L24
+Macromolecule #26: 60S ribosomal protein L23a
+Macromolecule #27: 60S ribosomal protein L26
+Macromolecule #28: 60S ribosomal protein L27
+Macromolecule #29: 60S ribosomal protein L27a
+Macromolecule #30: 60S ribosomal protein L29
+Macromolecule #31: 60S ribosomal protein L30
+Macromolecule #32: 60S ribosomal protein L31
+Macromolecule #33: 60S ribosomal protein L32
+Macromolecule #34: 60S ribosomal protein L35a
+Macromolecule #35: 60S ribosomal protein L34
+Macromolecule #36: Ribosomal protein uL29
+Macromolecule #37: 60S ribosomal protein L36
+Macromolecule #38: 60S ribosomal protein L37
+Macromolecule #39: 60S ribosomal protein L38
+Macromolecule #40: 60S ribosomal protein L39
+Macromolecule #41: Ubiquitin-60S ribosomal protein L40
+Macromolecule #42: 60S ribosomal protein L36a
+Macromolecule #43: 60S ribosomal protein L37a
+Macromolecule #44: 60S ribosomal protein L28
+Macromolecule #45: Eukaryotic translation initiation factor 6
+Macromolecule #46: MAGNESIUM ION
+Macromolecule #47: POTASSIUM ION
+Macromolecule #48: SODIUM ION
+Macromolecule #49: (3beta)-O~3~-[(2R)-2,6-dihydroxy-2-(2-methoxy-2-oxoethyl)-6-methy...
+Macromolecule #50: SPERMINE
+Macromolecule #51: 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID
+Macromolecule #52: ZINC ION
+Macromolecule #53: water
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2.0 nm Details: Grid was functionalised with 1-pyrenemethylamine. See methods section in published article. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 30 seconds 'wait' time. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 9973 / Average exposure time: 2.2 sec. / Average electron dose: 51.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 1147785 |
|---|---|
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID:  6ek0 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 3) |
| Final 3D classification | Number classes: 3 / Software - Name: cryoSPARC (ver. 3.1) Details: Used to filter out very few bad particles, mostly due to ice contamination. |
| Final angle assignment | Type: OTHER / Software - Name: RELION (ver. 3.1) |
| Final reconstruction | Number classes used: 1 / Resolution.type: BY AUTHOR / Resolution: 1.67 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1) / Number images used: 1088709 |
FSC plot (resolution estimation) |  |
 Movie
Movie Controller
Controller

















 Z
Z Y
Y X
X