[English] 日本語
 Yorodumi
Yorodumi- EMDB-14124: Structure of Bacillus pseudofirmus Mrp antiporter complex, monomer -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Bacillus pseudofirmus Mrp antiporter complex, monomer | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information: / monoatomic ion transmembrane transporter activity /  antiporter activity / inorganic cation transmembrane transport / monoatomic cation transmembrane transporter activity / sodium ion transport / antiporter activity / inorganic cation transmembrane transport / monoatomic cation transmembrane transporter activity / sodium ion transport /  NADH dehydrogenase (ubiquinone) activity / ATP synthesis coupled electron transport / monoatomic ion transport / proton transmembrane transport ...: / monoatomic ion transmembrane transporter activity / NADH dehydrogenase (ubiquinone) activity / ATP synthesis coupled electron transport / monoatomic ion transport / proton transmembrane transport ...: / monoatomic ion transmembrane transporter activity /  antiporter activity / inorganic cation transmembrane transport / monoatomic cation transmembrane transporter activity / sodium ion transport / antiporter activity / inorganic cation transmembrane transport / monoatomic cation transmembrane transporter activity / sodium ion transport /  NADH dehydrogenase (ubiquinone) activity / ATP synthesis coupled electron transport / monoatomic ion transport / proton transmembrane transport / membrane => GO:0016020 / NADH dehydrogenase (ubiquinone) activity / ATP synthesis coupled electron transport / monoatomic ion transport / proton transmembrane transport / membrane => GO:0016020 /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Alkalihalobacillus pseudofirmus (bacteria) / Alkalihalobacillus pseudofirmus (bacteria) /   Alkalihalophilus pseudofirmus (bacteria) Alkalihalophilus pseudofirmus (bacteria) | |||||||||
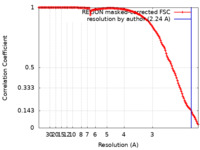
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.24 Å cryo EM / Resolution: 2.24 Å | |||||||||
 Authors Authors | Lee Y | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Ion transfer mechanisms in Mrp-type antiporters from high resolution cryoEM and molecular dynamics simulations. Authors: Yongchan Lee / Outi Haapanen / Anton Altmeyer / Werner Kühlbrandt / Vivek Sharma / Volker Zickermann /    Abstract: Multiple resistance and pH adaptation (Mrp) cation/proton antiporters are essential for growth of a variety of halophilic and alkaliphilic bacteria under stress conditions. Mrp-type antiporters are ...Multiple resistance and pH adaptation (Mrp) cation/proton antiporters are essential for growth of a variety of halophilic and alkaliphilic bacteria under stress conditions. Mrp-type antiporters are closely related to the membrane domain of respiratory complex I. We determined the structure of the Mrp antiporter from Bacillus pseudofirmus by electron cryo-microscopy at 2.2 Å resolution. The structure resolves more than 99% of the sidechains of the seven membrane subunits MrpA to MrpG plus 360 water molecules, including ~70 in putative ion translocation pathways. Molecular dynamics simulations based on the high-resolution structure revealed details of the antiport mechanism. We find that switching the position of a histidine residue between three hydrated pathways in the MrpA subunit is critical for proton transfer that drives gated trans-membrane sodium translocation. Several lines of evidence indicate that the same histidine-switch mechanism operates in respiratory complex I. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14124.map.gz emd_14124.map.gz | 10.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14124-v30.xml emd-14124-v30.xml emd-14124.xml emd-14124.xml | 21.9 KB 21.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14124_fsc.xml emd_14124_fsc.xml | 14 KB | Display |  FSC data file FSC data file |
| Images |  emd_14124.png emd_14124.png | 91.2 KB | ||
| Masks |  emd_14124_msk_1.map emd_14124_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Others |  emd_14124_half_map_1.map.gz emd_14124_half_map_1.map.gz emd_14124_half_map_2.map.gz emd_14124_half_map_2.map.gz | 194.7 MB 194.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14124 http://ftp.pdbj.org/pub/emdb/structures/EMD-14124 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14124 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14124 | HTTPS FTP |
-Related structure data
| Related structure data |  7qruMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14124.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14124.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07136 Å | ||||||||||||||||||||||||||||||||||||
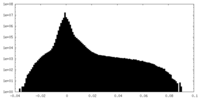
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14124_msk_1.map emd_14124_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
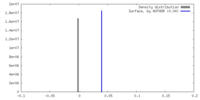
| Density Histograms |
-Half map: #2
| File | emd_14124_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
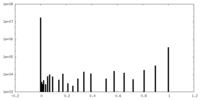
| Density Histograms |
-Half map: #1
| File | emd_14124_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
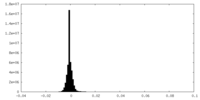
| Density Histograms |
- Sample components
Sample components
+Entire : MrpABCDEFG complex
+Supramolecule #1: MrpABCDEFG complex
+Macromolecule #1: Na+/H+ antiporter subunit D
+Macromolecule #2: Na+/H+ antiporter subunit A
+Macromolecule #3: Na(+)/H(+) antiporter subunit B
+Macromolecule #4: Na(+)/H(+) antiporter subunit C
+Macromolecule #5: Na+/H+ antiporter subunit E
+Macromolecule #6: Na(+)/H(+) antiporter subunit F
+Macromolecule #7: Na+/H+ antiporter subunit G1
+Macromolecule #8: 1,2-Distearoyl-sn-glycerophosphoethanolamine
+Macromolecule #9: water
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)