+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13700 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
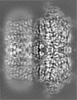
| Title | Ca2+ bound Drosophila Slo channel | |||||||||
 Map data Map data | Sharpened by Phenix.Autosharpen | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationSperm Motility And Taxes / negative regulation of neuromuscular synaptic transmission / large conductance calcium-activated potassium channel activity / regulation of synaptic assembly at neuromuscular junction / male courtship behavior, veined wing generated song production /  calcium-activated potassium channel activity / circadian behavior / monoatomic ion channel complex / potassium ion transmembrane transport / potassium ion transport ...Sperm Motility And Taxes / negative regulation of neuromuscular synaptic transmission / large conductance calcium-activated potassium channel activity / regulation of synaptic assembly at neuromuscular junction / male courtship behavior, veined wing generated song production / calcium-activated potassium channel activity / circadian behavior / monoatomic ion channel complex / potassium ion transmembrane transport / potassium ion transport ...Sperm Motility And Taxes / negative regulation of neuromuscular synaptic transmission / large conductance calcium-activated potassium channel activity / regulation of synaptic assembly at neuromuscular junction / male courtship behavior, veined wing generated song production /  calcium-activated potassium channel activity / circadian behavior / monoatomic ion channel complex / potassium ion transmembrane transport / potassium ion transport / calcium-activated potassium channel activity / circadian behavior / monoatomic ion channel complex / potassium ion transmembrane transport / potassium ion transport /  circadian rhythm / circadian rhythm /  postsynaptic membrane / neuron projection / response to xenobiotic stimulus / neuronal cell body / postsynaptic membrane / neuron projection / response to xenobiotic stimulus / neuronal cell body /  membrane / membrane /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.38 Å cryo EM / Resolution: 2.38 Å | |||||||||
 Authors Authors | Raisch T / Brockmann A / Ebbinghaus-Kintscher U / Freigang J / Gutbrod O / Kubicek J / Maertens B / Hofnagel O / Raunser S | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Small molecule modulation of the Drosophila Slo channel elucidated by cryo-EM. Authors: Tobias Raisch / Andreas Brockmann / Ulrich Ebbinghaus-Kintscher / Jörg Freigang / Oliver Gutbrod / Jan Kubicek / Barbara Maertens / Oliver Hofnagel / Stefan Raunser /  Abstract: Slowpoke (Slo) potassium channels display extraordinarily high conductance, are synergistically activated by a positive transmembrane potential and high intracellular Ca concentrations and are ...Slowpoke (Slo) potassium channels display extraordinarily high conductance, are synergistically activated by a positive transmembrane potential and high intracellular Ca concentrations and are important targets for insecticides and antiparasitic drugs. However, it is unknown how these compounds modulate ion translocation and whether there are insect-specific binding pockets. Here, we report structures of Drosophila Slo in the Ca-bound and Ca-free form and in complex with the fungal neurotoxin verruculogen and the anthelmintic drug emodepside. Whereas the architecture and gating mechanism of Slo channels are conserved, potential insect-specific binding pockets exist. Verruculogen inhibits K transport by blocking the Ca-induced activation signal and precludes K from entering the selectivity filter. Emodepside decreases the conductance by suboptimal K coordination and uncouples ion gating from Ca and voltage sensing. Our results expand the mechanistic understanding of Slo regulation and lay the foundation for the rational design of regulators of Slo and other voltage-gated ion channels. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13700.map.gz emd_13700.map.gz | 65 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13700-v30.xml emd-13700-v30.xml emd-13700.xml emd-13700.xml | 22.5 KB 22.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13700_fsc.xml emd_13700_fsc.xml | 17.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_13700.png emd_13700.png | 229.2 KB | ||
| Others |  emd_13700_additional_1.map.gz emd_13700_additional_1.map.gz emd_13700_additional_2.map.gz emd_13700_additional_2.map.gz emd_13700_half_map_1.map.gz emd_13700_half_map_1.map.gz emd_13700_half_map_2.map.gz emd_13700_half_map_2.map.gz | 24.8 MB 32.7 MB 164.8 MB 164.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13700 http://ftp.pdbj.org/pub/emdb/structures/EMD-13700 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13700 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13700 | HTTPS FTP |
-Related structure data
| Related structure data |  7pxeMC  7pxfC  7pxgC  7pxhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13700.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13700.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened by Phenix.Autosharpen | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Map sharpened by SPHIRE
| File | emd_13700_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map sharpened by SPHIRE | ||||||||||||
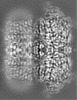
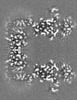
| Projections & Slices |
| ||||||||||||
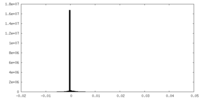
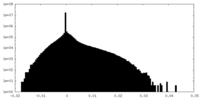
| Density Histograms |
-Additional map: Density-modified using Phenix.Reslove
| File | emd_13700_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Density-modified using Phenix.Reslove | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_13700_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_13700_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Slo tetramer
| Entire | Name: Slo tetramer |
|---|---|
| Components |
|
-Supramolecule #1: Slo tetramer
| Supramolecule | Name: Slo tetramer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) |
| Recombinant expression | Organism:   Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
-Macromolecule #1: Isoform J of Calcium-activated potassium channel slowpoke
| Macromolecule | Name: Isoform J of Calcium-activated potassium channel slowpoke type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) |
| Molecular weight | Theoretical: 130.919094 KDa |
| Recombinant expression | Organism:   Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASGLIDTNF SSTLANGMSG CDQSTVESLA DDPTDSPFDA DDCLKVRKYW CFLLSSIFTF LAGLLVVLLW RAFAFVCCRK EPDLGPNDP KQKEQKASRN KQEFEGTFMT EAKDWAGELI SGQTTTGRIL VVLVFILSIA SLIIYFVDAS SEEVERCQKW S NNITQQID ...String: MASGLIDTNF SSTLANGMSG CDQSTVESLA DDPTDSPFDA DDCLKVRKYW CFLLSSIFTF LAGLLVVLLW RAFAFVCCRK EPDLGPNDP KQKEQKASRN KQEFEGTFMT EAKDWAGELI SGQTTTGRIL VVLVFILSIA SLIIYFVDAS SEEVERCQKW S NNITQQID LAFNIFFMVY FFIRFIAASD KLWFMLEMYS FVDYFTIPPS FVSIYLDRTW IGLRFLRALR LMTVPDILQY LN VLKTSSS IRLAQLVSIF ISVWLTAAGI IHLLENSGDP LDFDNAHRLS YWTCVYFLIV TMSTVGYGDV YCETVLGRTF LVF FLLVGL AIFASCIPEI IDLIGTRAKY GGTLKNEKGR RHIVVCGHIT YESVSHFLKD FLHEDREDVD VEVVFLHRKP PDLE LEGLF KRHFTTVEFF QGTIMNPIDL QRVKVHEADA CLVLANKYCQ DPDAEDAANI MRVISIKNYS DDIRVIIQLM QYHNK AYLL NIPSWDWKQG DDVICLAELK LGFIAQSCLA PGFSTMMANL FAMRSFKTSP DMQSWTNDYL RGTGMEMYTE TLSPTF IGI PFAQATELCF SKLKLLLLAI EIKGAEEGAD SKISINPRGA KIQANTQGFF IAQSADEVKR AWFYCKACHE DIKDETL IK KCKCKNLATF RKGVRAVQMV GRASDITRDR EDTNLLNRNV RRPNGTGNGT GGMHHMNNTA AAAAAAAAAG KQVNKVKP T VNVSRQVEGQ VISPSQYNRP TSRSSGTGTQ NQNGGVSLPA GIADDQSKDF DFEKTEMKYD STGMFHWSPA KSLEDCILD RNQAAMTVLN GHVVVCLFAD PDSPLIGLRN LVMPLRASNF HYHELKHVVI VGSVDYIRRE WKMLQNLPKI SVLNGSPLSR ADLRAVNVN LCDMCCILSA KVPSNDDPTL ADKEAILASL NIKAMTFDDT IGVLSQRGPE FDNLSATAGS PIVLQRRGSV Y GANVPMIT ELVNDGNVQF LDQDDDDDPD TELYLTQPFA CGTAFAVSVL DSLMSTTYFN QNALTLIRSL ITGGATPELE LI LAEGAGL RGGYSTVESL SNRDRCRVGQ ISLYDGPLAQ FGECGKYGDL FVAALKSYGM LCIGLYRFRD TSSSCDASSK RYV ITNPPD DFSLLPTDQV FVLMQFDPGL EYKPPAVRAP AGGRGTNTQG SGVGGGGSNK DDNS |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 8 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #4: (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-...
| Macromolecule | Name: (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSAN-1-AMINIUM 4-OXIDE type: ligand / ID: 4 / Number of copies: 16 / Formula: 6PL |
|---|---|
| Molecular weight | Theoretical: 763.1 Da |
| Chemical component information |  ChemComp-6PL: |
-Macromolecule #5: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 5 / Number of copies: 4 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #6: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 6 / Number of copies: 4 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Macromolecule #7: water
| Macromolecule | Name: water / type: ligand / ID: 7 / Number of copies: 136 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 74.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X