1Z19
 
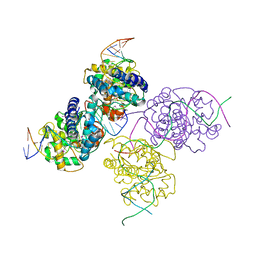 | | Crystal structure of a lambda integrase(75-356) dimer bound to a COC' core site | | Descriptor: | 33-MER, 5'-D(*CP*TP*CP*GP*TP*TP*CP*AP*GP*CP*TP*TP*TP*TP*TP*T)-3', 5'-D(P*TP*TP*TP*AP*TP*AP*CP*TP*AP*AP*GP*TP*TP*GP*GP*CP*AP*TP*TP*A)-3', ... | | Authors: | Biswas, T, Aihara, H, Radman-Livaja, M, Filman, D, Landy, A, Ellenberger, T. | | Deposit date: | 2005-03-03 | | Release date: | 2005-06-28 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | A structural basis for allosteric control of DNA recombination by lambda integrase.
Nature, 435, 2005
|
|
1Z1B
 
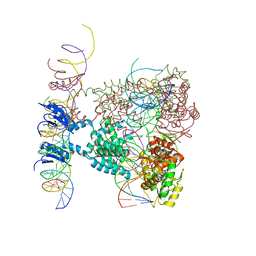 | | Crystal structure of a lambda integrase dimer bound to a COC' core site | | Descriptor: | 26-MER DNA, 29-MER DNA, 5'-D(*CP*T*CP*GP*TP*TP*CP*AP*GP*CP*TP*TP*TP*TP*TP*T)-3', ... | | Authors: | Biswas, T, Aihara, H, Radman-Livaja, M, Filman, D, Landy, A, Ellenberger, T. | | Deposit date: | 2005-03-03 | | Release date: | 2005-06-28 | | Last modified: | 2022-04-13 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | A structural basis for allosteric control of DNA recombination by lambda integrase.
Nature, 435, 2005
|
|
1Z1G
 
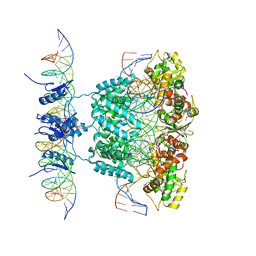 | | Crystal structure of a lambda integrase tetramer bound to a Holliday junction | | Descriptor: | 25-MER, 29-MER, 5'-D(*AP*CP*AP*GP*GP*TP*CP*AP*CP*TP*AP*TP*CP*AP*GP*TP*CP*AP*AP*AP*AP*TP*AP*CP*C)-3', ... | | Authors: | Biswas, T, Aihara, H, Radman-Livaja, M, Filman, D, Landy, A, Ellenberger, T. | | Deposit date: | 2005-03-03 | | Release date: | 2005-06-28 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (4.4 Å) | | Cite: | A structural basis for allosteric control of DNA recombination by lambda integrase.
Nature, 435, 2005
|
|
3K6Z
 
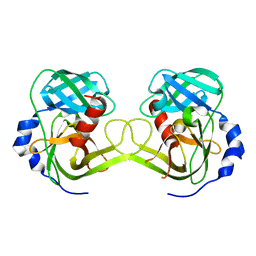 | | Crystal structure of Rv3671c protease, inactive form | | Descriptor: | POSSIBLE MEMBRANE-ASSOCIATED SERINE PROTEASE | | Authors: | Biswas, T, Small, J, Vandal, O, Ehrt, S, Tsodikov, O.V. | | Deposit date: | 2009-10-10 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural insight into serine protease Rv3671c that Protects M. tuberculosis from oxidative and acidic stress.
Structure, 18, 2010
|
|
3K6Y
 
 | | Crystal structure of Rv3671c protease from M. tuberculosis, active form | | Descriptor: | POSSIBLE MEMBRANE-ASSOCIATED SERINE PROTEASE | | Authors: | Biswas, T, Small, J, Vandal, O, Ehrt, S, Tsodikov, O.V. | | Deposit date: | 2009-10-10 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Structural insight into serine protease Rv3671c that Protects M. tuberculosis from oxidative and acidic stress.
Structure, 18, 2010
|
|
3LT3
 
 | | Crystal structure of Rv3671c from M. tuberculosis H37Rv, Ser343Ala mutant, inactive form | | Descriptor: | POSSIBLE MEMBRANE-ASSOCIATED SERINE PROTEASE | | Authors: | Biswas, T, Small, J, Vandal, O, Ehrt, S, Tsodikov, O.V. | | Deposit date: | 2010-02-14 | | Release date: | 2010-11-03 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural insight into serine protease Rv3671c that Protects M. tuberculosis from oxidative and acidic stress.
Structure, 18, 2010
|
|
3I6B
 
 | |
3ITW
 
 | |
2R5U
 
 | |
3U9F
 | |
3U9B
 
 | | Structure of apo-CATI | | Descriptor: | Chloramphenicol acetyltransferase | | Authors: | Biswas, T, Garneau-Tsodikova, S, Tsodikov, O.V. | | Deposit date: | 2011-10-18 | | Release date: | 2012-02-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | The structural basis for substrate versatility of chloramphenicol acetyltransferase CAT(I).
Protein Sci., 21, 2012
|
|
3HYC
 
 | |
3R1K
 
 | | Crystal structure of acetyltransferase Eis from Mycobacterium tuberculosis H37Rv in complex with CoA and an acetamide moiety | | Descriptor: | ACETAMIDE, COENZYME A, Enhanced intracellular survival protein | | Authors: | Biswas, T, Chen, W, Garneau-Tsodikova, S, Tsodikov, O.V. | | Deposit date: | 2011-03-10 | | Release date: | 2011-06-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Unusual regioversatility of acetyltransferase Eis, a cause of drug resistance in XDR-TB.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
2R8Z
 
 | | Crystal structure of YrbI phosphatase from Escherichia coli in complex with a phosphate and a calcium ion | | Descriptor: | 3-deoxy-D-manno-octulosonate 8-phosphate phosphatase, CALCIUM ION, PHOSPHATE ION | | Authors: | Tsodikov, O.V, Aggarwal, P, Rubin, J.R, Stuckey, J.A, Woodard, R.W, Biswas, T. | | Deposit date: | 2007-09-11 | | Release date: | 2008-09-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Tail of KdsC: CONFORMATIONAL CHANGES CONTROL THE ACTIVITY OF A HALOACID DEHALOGENASE SUPERFAMILY PHOSPHATASE.
J.Biol.Chem., 284, 2009
|
|
2R8X
 | | Crystal structure of YrbI phosphatase from Escherichia coli | | Descriptor: | 3-deoxy-D-manno-octulosonate 8-phosphate phosphatase, CHLORIDE ION | | Authors: | Tsodikov, O.V, Aggarwal, P, Rubin, J.R, Stuckey, J.A, Woodard, R.W, Biswas, T. | | Deposit date: | 2007-09-11 | | Release date: | 2008-09-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Tail of KdsC: CONFORMATIONAL CHANGES CONTROL THE ACTIVITY OF A HALOACID DEHALOGENASE SUPERFAMILY PHOSPHATASE.
J.Biol.Chem., 284, 2009
|
|
2R8Y
 | | Crystal structure of YrbI phosphatase from Escherichia coli in a complex with Ca | | Descriptor: | CALCIUM ION, CHLORIDE ION, YrbI from Escherichia coli | | Authors: | Tsodikov, O.V, Aggarwal, P, Rubin, J.R, Stuckey, J.A, Woodard, R.W, Biswas, T. | | Deposit date: | 2007-09-11 | | Release date: | 2008-09-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | The Tail of KdsC: CONFORMATIONAL CHANGES CONTROL THE ACTIVITY OF A HALOACID DEHALOGENASE SUPERFAMILY PHOSPHATASE.
J.Biol.Chem., 284, 2009
|
|
2R8E
 | | Crystal structure of YrbI from Escherichia coli in complex with Mg | | Descriptor: | 3-deoxy-D-manno-octulosonate 8-phosphate phosphatase, CHLORIDE ION, MAGNESIUM ION | | Authors: | Tsodikov, O.V, Aggarwal, P, Rubin, J.R, Stuckey, J.A, Woodard, R, Biswas, T. | | Deposit date: | 2007-09-10 | | Release date: | 2008-09-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The Tail of KdsC: CONFORMATIONAL CHANGES CONTROL THE ACTIVITY OF A HALOACID DEHALOGENASE SUPERFAMILY PHOSPHATASE.
J.Biol.Chem., 284, 2009
|
|
6B0U
 
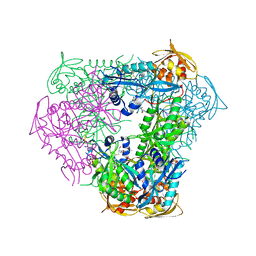 | | Crystal structure of acetyltransferase Eis from Mycobacterium tuberculosis in complex with a Lys-containing peptide | | Descriptor: | COENZYME A, N-acetyltransferase Eis, Synthetic peptide ATKAPAKKA | | Authors: | Biswas, T, Pang, A.H, Garneau-Tsodikova, S, Tsodikov, O.V. | | Deposit date: | 2017-09-15 | | Release date: | 2018-01-31 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Acetylation by Eis and Deacetylation by Rv1151c of Mycobacterium tuberculosis HupB: Biochemical and Structural Insight.
Biochemistry, 57, 2018
|
|
4I9G
 
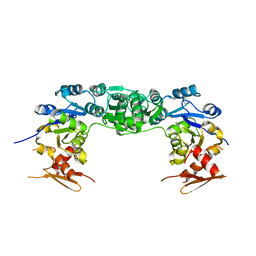 | | Crystal structure of glycerol phosphate phosphatase Rv1692 from Mycobacterium tuberculosis in complex with magnesium | | Descriptor: | Glycerol 3-phosphate phosphatase, MAGNESIUM ION | | Authors: | Biswas, T, Larrouy-Maumus, G, de Carvalho, L.P, Tsodikov, O.V. | | Deposit date: | 2012-12-05 | | Release date: | 2013-07-10 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Discovery of a glycerol 3-phosphate phosphatase reveals glycerophospholipid polar head recycling in Mycobacterium tuberculosis.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4I9F
 | | Crystal structure of glycerol phosphate phosphatase Rv1692 from Mycobacterium tuberculosis in complex with calcium | | Descriptor: | CALCIUM ION, CHLORIDE ION, Glycerol 3-phosphate phosphatase | | Authors: | Biswas, T, Larrouy-Maumus, G, de Carvalho, L.P, Tsodikov, O.V. | | Deposit date: | 2012-12-05 | | Release date: | 2013-07-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | Discovery of a glycerol 3-phosphate phosphatase reveals glycerophospholipid polar head recycling in Mycobacterium tuberculosis.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4IJJ
 
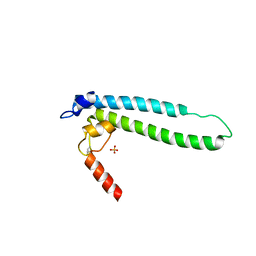 | | Structure of transcription factor DksA2 from Pseudomonas aeruginosa | | Descriptor: | Putative C4-type zinc finger protein, DksA/TraR family, SULFATE ION | | Authors: | Biswas, T, Furman, R, Artsimovitch, I, Tsodikov, O.V. | | Deposit date: | 2012-12-21 | | Release date: | 2013-02-27 | | Last modified: | 2013-03-27 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | DksA2, a zinc-independent structural analog of the transcription factor DksA.
Febs Lett., 587, 2013
|
|
4JD6
 
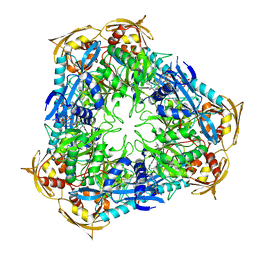 | | Crystal structure of Mycobacterium tuberculosis Eis in complex with coenzyme A and tobramycin | | Descriptor: | COENZYME A, Enhanced intracellular survival protein, TOBRAMYCIN | | Authors: | Biswas, T, Chen, W, Garneau-Tsodikova, S, Tsodikov, O.V. | | Deposit date: | 2013-02-23 | | Release date: | 2013-10-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Chemical and structural insights into the regioversatility of the aminoglycoside acetyltransferase eis.
Chembiochem, 14, 2013
|
|
4Z74
 
 | |
4Z71
 
 | |
4Z72
 
 | | Crystal structure of inorganic pyrophosphatase from Mycobacterium tuberculosis in complex with two phosphate ions | | Descriptor: | CALCIUM ION, Inorganic pyrophosphatase, PHOSPHATE ION | | Authors: | Pratt, A.C, Biswas, T, Barnard-Britson, S, Tsodikov, O.V. | | Deposit date: | 2015-04-06 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.352 Å) | | Cite: | Structural and computational dissection of the catalytic mechanism of the inorganic pyrophosphatase from Mycobacterium tuberculosis.
J.Struct.Biol., 192, 2015
|
|
