4WF2
 
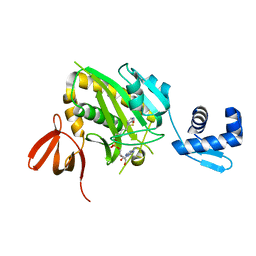 | | Structure of E. coli BirA G142A bound to biotinol-5'-AMP | | Descriptor: | ((2R,3S,4R,5R)-5-(6-AMINO-9H-PURIN-9-YL)-3,4-DIHYDROXY-TETRAHYDROFURAN-2-YL)METHYL 5-((3AS,4S,6AR)-2-OXO-HEXAHYDRO-1H-THIENO[3,4-D]IMIDAZOL-4-YL)PENTYL HYDROGEN PHOSPHATE, Bifunctional ligase/repressor BirA | | Authors: | Eginton, C, Beckett, D, Wade, H. | | Deposit date: | 2014-09-11 | | Release date: | 2014-10-22 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Allosteric Coupling via Distant Disorder-to-Order Transitions.
J.Mol.Biol., 427, 2015
|
|
3Q1M
 
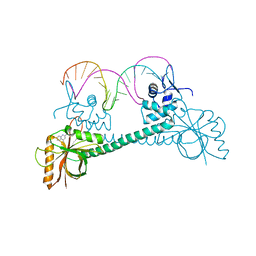 | | Crystal Structure of BmrR Dimer bound to DNA and the ligand 4-amino-quinaldine | | Descriptor: | 2-methylquinolin-4-amine, 23 bp promoter DNA, Multidrug-efflux transporter 1 regulator | | Authors: | Bachas, S, Eginton, C, Gunio, G, Wade, H. | | Deposit date: | 2010-12-17 | | Release date: | 2011-06-15 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural contributions to multidrug recognition in the multidrug resistance (MDR) gene regulator, BmrR.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3Q2Y
 
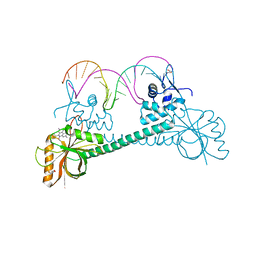 | | Crystal Structure of BmrR bound to ethidium | | Descriptor: | 23 bp promoter, ETHIDIUM, GLYCEROL, ... | | Authors: | Bachas, S, Eginton, C, Gunio, G, Wade, H. | | Deposit date: | 2010-12-20 | | Release date: | 2011-06-15 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural contributions to multidrug recognition in the multidrug resistance (MDR) gene regulator, BmrR.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3Q3D
 
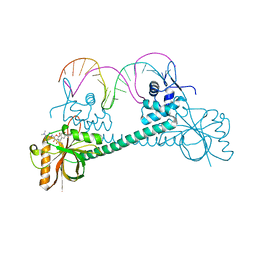 | | Crystal structure of BmrR bound to puromycin | | Descriptor: | BmrR promoter DNA, Multidrug-efflux transporter 1 regulator, PUROMYCIN | | Authors: | Bachas, S, Eginton, C, Gunio, G, Wade, H. | | Deposit date: | 2010-12-21 | | Release date: | 2011-06-15 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | Structural contributions to multidrug recognition in the multidrug resistance (MDR) gene regulator, BmrR.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3Q5P
 
 | | Crystal structure of BmrR bound to Tetracycline | | Descriptor: | 23 bp BmrR promoter DNA, GLYCEROL, Multidrug-efflux transporter 1 regulator, ... | | Authors: | Bachas, S, Eginton, C, Gunio, G, Wade, H. | | Deposit date: | 2010-12-29 | | Release date: | 2011-06-15 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.942 Å) | | Cite: | Structural contributions to multidrug recognition in the multidrug resistance (MDR) gene regulator, BmrR.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3Q5R
 
 | | Crystal structure of BmrR bound to Kanamycin | | Descriptor: | 23 bp promoter DNA, KANAMYCIN A, Multidrug-efflux transporter 1 regulator | | Authors: | Bachas, S, Eginton, C, Gunio, G, Wade, H. | | Deposit date: | 2010-12-29 | | Release date: | 2011-07-20 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Structural contributions to multidrug recognition in the multidrug resistance (MDR) gene regulator, BmrR.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3Q5S
 
 | | Crystal structure of BmrR bound to Acetylcholine | | Descriptor: | 23 bp promoter DNA, ACETYLCHOLINE, GLYCEROL, ... | | Authors: | Bachas, S, Eginton, C, Gunio, G, Wade, H. | | Deposit date: | 2010-12-29 | | Release date: | 2011-06-15 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural contributions to multidrug recognition in the multidrug resistance (MDR) gene regulator, BmrR.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
5KAX
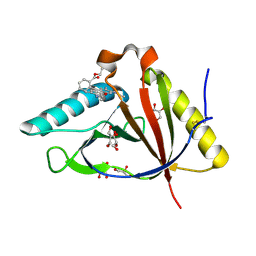 
 | | The structure of CTR107 protein bound to RHODAMINE 6G | | Descriptor: | CTR107 protein, GLYCEROL, RHODAMINE 6G | | Authors: | Moreno, A, Wade, H. | | Deposit date: | 2016-06-02 | | Release date: | 2016-08-24 | | Last modified: | 2019-11-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Solution Binding and Structural Analyses Reveal Potential Multidrug Resistance Functions for SAV2435 and CTR107 and Other GyrI-like Proteins.
Biochemistry, 55, 2016
|
|
5KAU
 
 | | The structure of SAV2435 bound to RHODAMINE 6G | | Descriptor: | GLYCEROL, RHODAMINE 6G, SA2223 protein | | Authors: | Moreno, A, Wade, H. | | Deposit date: | 2016-06-02 | | Release date: | 2016-08-24 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Solution Binding and Structural Analyses Reveal Potential Multidrug Resistance Functions for SAV2435 and CTR107 and Other GyrI-like Proteins.
Biochemistry, 55, 2016
|
|
5KAW
 
 | |
5KAV
 
 | | The structure of SAV2435 | | Descriptor: | GLYCEROL, SA2223 protein | | Authors: | Moreno, A, Wade, H. | | Deposit date: | 2016-06-02 | | Release date: | 2016-08-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Solution Binding and Structural Analyses Reveal Potential Multidrug Resistance Functions for SAV2435 and CTR107 and Other GyrI-like Proteins.
Biochemistry, 55, 2016
|
|
5KAT
 
 | | The structure of SAV2435 bound to TETRAPHENYLPHOSPHONIUM | | Descriptor: | GLYCEROL, SA2223 protein, TETRAPHENYLPHOSPHONIUM | | Authors: | Moreno, A, Wade, H. | | Deposit date: | 2016-06-02 | | Release date: | 2016-08-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.101 Å) | | Cite: | Solution Binding and Structural Analyses Reveal Potential Multidrug Resistance Functions for SAV2435 and CTR107 and Other GyrI-like Proteins.
Biochemistry, 55, 2016
|
|
5KCB
 
 | | The structure of SAV2435 bound to ethidium bromide | | Descriptor: | ETHIDIUM, SA2223 protein, SULFATE ION | | Authors: | Moreno, A, Wade, H. | | Deposit date: | 2016-06-06 | | Release date: | 2016-08-24 | | Last modified: | 2019-11-27 | | Method: | X-RAY DIFFRACTION (2.101 Å) | | Cite: | Solution Binding and Structural Analyses Reveal Potential Multidrug Resistance Functions for SAV2435 and CTR107 and Other GyrI-like Proteins.
Biochemistry, 55, 2016
|
|
1JM0
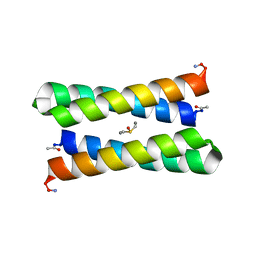 
 | | CRYSTAL STRUCTURE OF FOUR-HELIX BUNDLE MODEL | | Descriptor: | DIMETHYL SULFOXIDE, MANGANESE (II) ION, PROTEIN (FOUR-HELIX BUNDLE MODEL) | | Authors: | Di Costanzo, L, Geremia, S. | | Deposit date: | 2001-07-17 | | Release date: | 2002-01-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Toward the de novo design of a catalytically active helix bundle: a substrate-accessible carboxylate-bridged dinuclear metal center.
J.Am.Chem.Soc., 123, 2001
|
|
1JMB
 
 | | CRYSTAL STRUCTURE OF FOUR-HELIX BUNDLE MODEL | | Descriptor: | DIMETHYL SULFOXIDE, MANGANESE (II) ION, PROTEIN (FOUR-HELIX BUNDLE MODEL) | | Authors: | Di Costanzo, L, Geremia, S. | | Deposit date: | 2001-07-18 | | Release date: | 2002-01-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Toward the de novo design of a catalytically active helix bundle: a substrate-accessible carboxylate-bridged dinuclear metal center.
J.Am.Chem.Soc., 123, 2001
|
|
1LT1
 
 | |
2KIK
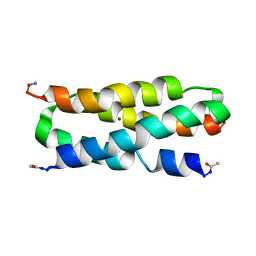 
 | |
1NVO
 
 | | Solution structure of a four-helix bundle model, apo-DF1 | | Descriptor: | Homodimeric Alpha2 Four-Helix Bundle | | Authors: | Maglio, O, Nastri, F, Pavone, V, Lombardi, A, DeGrado, W.F. | | Deposit date: | 2003-02-04 | | Release date: | 2003-03-25 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Preorganization of molecular binding sites in designed diiron proteins
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1OVV
 
 | |
1OVU
 
 | |
1OVR
 
 | | CRYSTAL STRUCTURE OF FOUR-HELIX BUNDLE MODEL di-Mn(II)-DF1-L13 | | Descriptor: | MANGANESE (II) ION, four-helix bundle model di-Mn(II)-DF1-L13 | | Authors: | Di Costanzo, L, Geremia, S. | | Deposit date: | 2003-03-27 | | Release date: | 2004-05-18 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | Response of a designed metalloprotein to changes in metal ion coordination, exogenous ligands, and active site volume determined by X-ray crystallography.
J.Am.Chem.Soc., 127, 2005
|
|
