6QAX
 
 | | P31-43 | | Descriptor: | LEU-GLY-GLN-GLN-GLN-PRO-PHE-PRO-PRO-GLN-GLN-PRO-TYR | | Authors: | Calvanese, L, D'Auria, G, Falcigno, F. | | Deposit date: | 2018-12-19 | | Release date: | 2019-02-27 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural insights on P31-43, a gliadin peptide able to promote an innate but not an adaptive response in celiac disease.
J.Pept.Sci., 25, 2019
|
|
7OB2
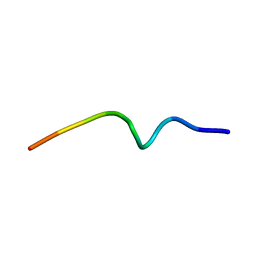 
 | | NMR structure of the antimicrobial RiLK1 peptide in SDS micelles | | Descriptor: | RiLK1 | | Authors: | Falcigno, L, D'Auria, G, Palmieri, G, Gogliettino, M, Agrillo, B. | | Deposit date: | 2021-04-20 | | Release date: | 2021-11-17 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Key Physicochemical Determinants in the Antimicrobial Peptide RiLK1 Promote Amphipathic Structures.
Int J Mol Sci, 22, 2021
|
|
2J5H
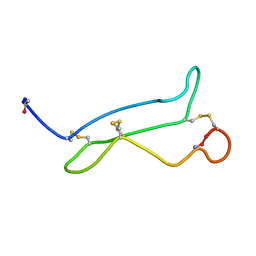 
 | | NMR analysis of mouse CRIPTO CFC domain | | Descriptor: | TERATOCARCINOMA-DERIVED GROWTH FACTOR | | Authors: | Calvanese, L, Saporito, A, Marasco, D, D'Auria, G, Minchiotti, G, Pedone, C, Paolillo, L, Falcigno, L, Ruvo, M. | | Deposit date: | 2006-09-18 | | Release date: | 2006-10-02 | | Last modified: | 2018-05-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure of mouse Cripto CFC domain and its inactive variant Trp107Ala.
J. Med. Chem., 49, 2006
|
|
6QB0
 
 | | 5675 | | Descriptor: | LEU-GLY-GLN-GLN-GLN-ALA-PHE-PRO-PRO-GLN-GLN-PRO-TYR | | Authors: | Calvanese, L, D'Auria, G, Falcigno, F. | | Deposit date: | 2018-12-20 | | Release date: | 2019-02-27 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural insights on P31-43, a gliadin peptide able to promote an innate but not an adaptive response in celiac disease.
J.Pept.Sci., 25, 2019
|
|
6QB1
 
 | | 5676 | | Descriptor: | LEU-GLY-GLN-GLN-GLN-PRO-ALA-PRO-PRO-GLN-GLN-PRO-TYR | | Authors: | Calvanese, L, D'Auria, G, Falcigno, F. | | Deposit date: | 2018-12-20 | | Release date: | 2019-02-27 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural insights on P31-43, a gliadin peptide able to promote an innate but not an adaptive response in celiac disease.
J.Pept.Sci., 25, 2019
|
|
2MGV
 
 | | NMR structure of PASTA domain of PonA2 from Mycobacterium tuberculosis | | Descriptor: | Bifunctional membrane-associated penicillin-binding protein 1A/1B ponA2 | | Authors: | Calvanese, L, Falcigno, L, Maglione, C, Marasco, D, Ruggiero, A, Squeglia, F, Berisio, R, D'Auria, G. | | Deposit date: | 2013-11-11 | | Release date: | 2013-12-25 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural and binding properties of the PASTA domain of PonA2, a key penicillin binding protein from Mycobacterium tuberculosis.
Biopolymers, 101, 2014
|
|
6S0M
 
 | |
6EY3
 
 | |
